Ku and DNA-dependent protein kinase dynamic conformations and assembly regulate DNA binding and the initial non-homologous end joining complex
- PMID: 19893054
- PMCID: PMC2801267
- DOI: 10.1074/jbc.M109.065615
Ku and DNA-dependent protein kinase dynamic conformations and assembly regulate DNA binding and the initial non-homologous end joining complex
Abstract
DNA double strand break (DSB) repair by non-homologous end joining (NHEJ) is initiated by DSB detection by Ku70/80 (Ku) and DNA-dependent protein kinase catalytic subunit (DNA-PKcs) recruitment, which promotes pathway progression through poorly defined mechanisms. Here, Ku and DNA-PKcs solution structures alone and in complex with DNA, defined by x-ray scattering, reveal major structural reorganizations that choreograph NHEJ initiation. The Ku80 C-terminal region forms a flexible arm that extends from the DNA-binding core to recruit and retain DNA-PKcs at DSBs. Furthermore, Ku- and DNA-promoted assembly of a DNA-PKcs dimer facilitates trans-autophosphorylation at the DSB. The resulting site-specific autophosphorylation induces a large conformational change that opens DNA-PKcs and promotes its release from DNA ends. These results show how protein and DNA interactions initiate large Ku and DNA-PKcs rearrangements to control DNA-PK biological functions as a macromolecular machine orchestrating assembly and disassembly of the initial NHEJ complex on DNA.
Figures
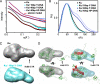



References
-
- Hall E. J., Giaccia A. J. (2006) Radiobiology for the Radiobiologist, 6th Ed., pp. 16–46, Lippincott Williams & Wilkins, Philadelphia
-
- Helleday T., Lo J., van Gent D. C., Engelward B. P. (2007) DNA Repair 6, 923–935 - PubMed
-
- Löbrich M., Jeggo P. A. (2007) Nat. Rev. Cancer 7, 861–869 - PubMed
-
- O'Driscoll M., Jeggo P. A. (2006) Nat. Rev. Genet 7, 45–54 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

