Re-examination of siRNA specificity questions role of PICH and Tao1 in the spindle checkpoint and identifies Mad2 as a sensitive target for small RNAs
- PMID: 19904549
- PMCID: PMC2846388
- DOI: 10.1007/s00412-009-0244-2
Re-examination of siRNA specificity questions role of PICH and Tao1 in the spindle checkpoint and identifies Mad2 as a sensitive target for small RNAs
Abstract
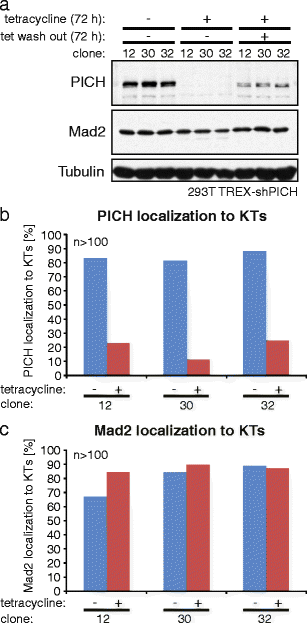
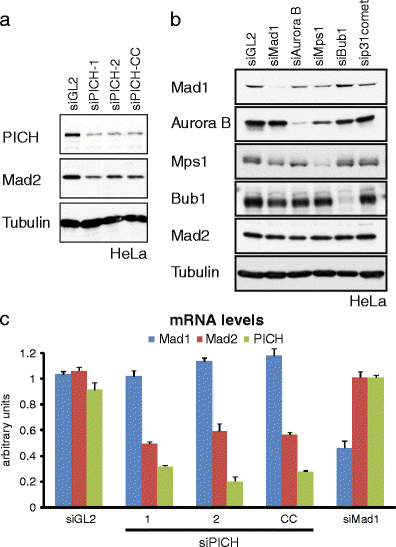
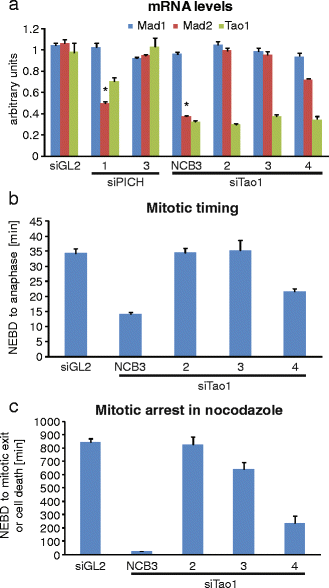
The DNA-dependent adenosine triphosphatase (ATPase) Plk1-interacting checkpoint helicase (PICH) has recently been implicated in spindle checkpoint (SAC) signaling (Baumann et al., Cell 128(1):101-114, 2007). Depletion of PICH by siRNA abolished the SAC and resulted in an apparently selective loss of Mad2 from kinetochores, suggesting a role for PICH in the regulation of the Mad1-Mad2 interaction. An apparent rescue of SAC functionality by overexpression of PICH in PICH-depleted cells initially seemed to confirm a role for PICH in the SAC. However, we have subsequently discovered that all PICH-directed siRNA oligonucleotides that abolish the SAC also reduce Mad2 mRNA and protein expression. This reduction is functionally significant, as PICH siRNA does not abolish SAC activity in a cell line that harbors a bacterial artificial chromosome driving the expression of murine Mad2. Moreover, we identified several siRNA duplexes that effectively deplete PICH but do not significantly affect SAC functionality or Mad2 abundance or localization. Finally, we discovered that the ability of overexpressed PICH to restore SAC activity in PICH-depleted cells depends on sequestration of the mitotic kinase Plk1 rather than ATPase activity of PICH, pointing to an underlying mechanism of "bypass suppression." In support of this view, depletion or inhibition of Plk1 also rescued SAC activity in cells harboring low levels of Mad2. This observation suggests that a reduction of Plk1 activity partially compensates for reduced Mad2 levels and argues that Plk1 normally reduces the strength of SAC signaling. Collectively, our results question the role of PICH in the SAC and instead identify Mad2 as a sensitive off target for small RNA duplexes. In support of the latter conclusion, our evidence suggests that an off-target effect on Mad2 may also contribute to explain the apparent role of the Tao1 kinase in SAC signaling.
Figures





References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

