Sculpting the bacterial cell
- PMID: 19906583
- PMCID: PMC4080913
- DOI: 10.1016/j.cub.2009.06.033
Sculpting the bacterial cell
Abstract
Prokaryotes come in a wide variety of shapes, determined largely by natural selection, physical constraints, and patterns of cell growth and division. Because of their relative simplicity, bacterial cells are excellent models for how genes and proteins can directly determine morphology. Recent advances in cytological methods for bacteria have shown that distinct cytoskeletal filaments composed of actin and tubulin homologs are important for guiding growth patterns of the cell wall in bacteria, and that the glycan strands that constitute the wall are generally perpendicular to the direction of growth. This cytoskeleton-directed cell wall patterning is strikingly reminiscent of how plant cell wall growth is regulated by microtubules. In rod-shaped bacilli, helical cables of actin-like MreB protein stretch along the cell length and orchestrate elongation of the cell wall, whereas the tubulin-like FtsZ protein directs formation of the division septum and the resulting cell poles. The overlap and interplay between these two systems and the peptidoglycan-synthesizing enzymes they recruit are the major driving forces of cylindrical shapes. Round cocci, on the other hand, have lost their MreB cables and instead must grow mainly via their division septum, giving them their characteristic round or ovoid shapes. Other bacteria that lack MreB homologs or even cell walls use distinct cytoskeletal systems to maintain their distinct shapes. Here I review what is known about the mechanisms that determine the shape of prokaryotic cells.
Figures
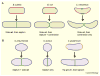
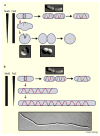
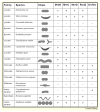
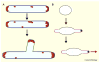
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

