JASPAR 2010: the greatly expanded open-access database of transcription factor binding profiles
- PMID: 19906716
- PMCID: PMC2808906
- DOI: 10.1093/nar/gkp950
JASPAR 2010: the greatly expanded open-access database of transcription factor binding profiles
Abstract
JASPAR (http://jaspar.genereg.net) is the leading open-access database of matrix profiles describing the DNA-binding patterns of transcription factors (TFs) and other proteins interacting with DNA in a sequence-specific manner. Its fourth major release is the largest expansion of the core database to date: the database now holds 457 non-redundant, curated profiles. The new entries include the first batch of profiles derived from ChIP-seq and ChIP-chip whole-genome binding experiments, and 177 yeast TF binding profiles. The introduction of a yeast division brings the convenience of JASPAR to an active research community. As binding models are refined by newer data, the JASPAR database now uses versioning of matrices: in this release, 12% of the older models were updated to improved versions. Classification of TF families has been improved by adopting a new DNA-binding domain nomenclature. A curated catalog of mammalian TFs is provided, extending the use of the JASPAR profiles to additional TFs belonging to the same structural family. The changes in the database set the system ready for more rapid acquisition of new high-throughput data sources. Additionally, three new special collections provide matrix profile data produced by recent alternative high-throughput approaches.
Figures
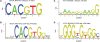
References
-
- Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods. 2008;5:621–628. - PubMed
-
- Carninci P, Sandelin A, Lenhard B, Katayama S, Shimokawa K, Ponjavic J, Semple CAM, Taylor MS, Engstrom PG, Frith MC, et al. Genome-wide analysis of mammalian promoter architecture and evolution. Nat. Genet. 2006;38:626–635. - PubMed
-
- Wasserman WW, Sandelin A. Applied bioinformatics for the identification of regulatory elements. Nat. Rev. Genet. 2004;5:276–287. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

