Accurate recognition of cis-regulatory motifs with the correct lengths in prokaryotic genomes
- PMID: 19906734
- PMCID: PMC2811016
- DOI: 10.1093/nar/gkp907
Accurate recognition of cis-regulatory motifs with the correct lengths in prokaryotic genomes
Abstract
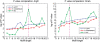
We present a new computational method for solving a classical problem, the identification problem of cis-regulatory motifs in a given set of promoter sequences, based on one key new idea. Instead of scoring candidate motifs individually like in all the existing motif-finding programs, our method scores groups of candidate motifs with similar sequences, called motif closures, using a P-value, which has substantially improved the prediction reliability over the existing methods. Our new P-value scoring scheme is sequence length independent, hence allowing direct comparisons among predicted motifs with different lengths on the same footing. We have implemented this method as a Motif Recognition Computer (MREC) program, and have extensively tested MREC on both simulated and biological data from prokaryotic genomes. Our test results indicate that MREC can accurately pick out the actual motif with the correct length as the best scoring candidate for the vast majority of the cases in our test set. We compared our prediction results with two motif-finding programs Cosmo and MEME, and found that MREC outperforms both programs across all the test cases by a large margin. The MREC program is available at http://csbl.bmb.uga.edu/~bingqiang/MREC1/.
Figures
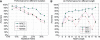
Similar articles
-
A new framework for identifying cis-regulatory motifs in prokaryotes.Nucleic Acids Res. 2011 Apr;39(7):e42. doi: 10.1093/nar/gkq948. Epub 2010 Dec 11. Nucleic Acids Res. 2011. PMID: 21149261 Free PMC article.
-
An integrative and applicable phylogenetic footprinting framework for cis-regulatory motifs identification in prokaryotic genomes.BMC Genomics. 2016 Aug 9;17:578. doi: 10.1186/s12864-016-2982-x. BMC Genomics. 2016. PMID: 27507169 Free PMC article.
-
An integrated toolkit for accurate prediction and analysis of cis-regulatory motifs at a genome scale.Bioinformatics. 2013 Sep 15;29(18):2261-8. doi: 10.1093/bioinformatics/btt397. Epub 2013 Jul 10. Bioinformatics. 2013. PMID: 23846744
-
Finding sequence motifs in prokaryotic genomes--a brief practical guide for a microbiologist.Brief Bioinform. 2009 Sep;10(5):525-36. doi: 10.1093/bib/bbp032. Epub 2009 Jun 24. Brief Bioinform. 2009. PMID: 19553402 Review.
-
The nature and dynamics of bacterial genomes.Science. 2006 Mar 24;311(5768):1730-3. doi: 10.1126/science.1119966. Science. 2006. PMID: 16556833 Review.
Cited by
-
Integrating genome sequence and structural data for statistical learning to predict transcription factor binding sites.Nucleic Acids Res. 2020 Dec 16;48(22):12604-12617. doi: 10.1093/nar/gkaa1134. Nucleic Acids Res. 2020. PMID: 33264415 Free PMC article.
-
Phylogenetic footprinting: a boost for microbial regulatory genomics.Protoplasma. 2012 Oct;249(4):901-7. doi: 10.1007/s00709-011-0351-9. Epub 2011 Nov 24. Protoplasma. 2012. PMID: 22113593 Review.
-
A new framework for identifying cis-regulatory motifs in prokaryotes.Nucleic Acids Res. 2011 Apr;39(7):e42. doi: 10.1093/nar/gkq948. Epub 2010 Dec 11. Nucleic Acids Res. 2011. PMID: 21149261 Free PMC article.
-
An integrative and applicable phylogenetic footprinting framework for cis-regulatory motifs identification in prokaryotic genomes.BMC Genomics. 2016 Aug 9;17:578. doi: 10.1186/s12864-016-2982-x. BMC Genomics. 2016. PMID: 27507169 Free PMC article.
References
-
- Stormo GD. DNA binding sites: representation and discovery. Bioinformatics. 2000;16:16–23. - PubMed
-
- Tompa M, Li N, Bailey TL, Church GM, De Moor B, Eskin E, Favorov AV, Frith MC, Fu Y, Kent WJ, et al. Assessing computational tools for the discovery of transcription factor binding sites. Nat. Biotechnol. 2005;23:137–144. - PubMed
-
- Thijs G, Marchal K, Lescot M, Rombauts S, De Moor B, Rouze P, Moreau Y. A Gibbs sampling method to detect overrepresented motifs in the upstream regions of coexpressed genes. J. Comput. Biol. 2002;9:447–464. - PubMed

