Genome-scale identification method applied to find cryptic aminoglycoside resistance genes in Pseudomonas aeruginosa
- PMID: 19907650
- PMCID: PMC2771283
- DOI: 10.1371/journal.pone.0006576
Genome-scale identification method applied to find cryptic aminoglycoside resistance genes in Pseudomonas aeruginosa
Abstract
Background: The ability of bacteria to rapidly evolve resistance to antibiotics is a critical public health problem. Resistance leads to increased disease severity and death rates, as well as imposes pressure towards the discovery and development of new antibiotic therapies. Improving understanding of the evolution and genetic basis of resistance is a fundamental goal in the field of microbiology.
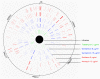
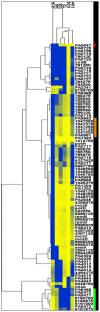
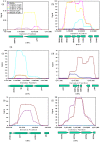
Results: We have applied a new genomic method, Scalar Analysis of Library Enrichments (SCALEs), to identify genomic regions that, given increased copy number, may lead to aminoglycoside resistance in Pseudomonas aeruginosa at the genome scale. We report the result of selections on highly representative genomic libraries for three different aminoglycoside antibiotics (amikacin, gentamicin, and tobramycin). At the genome-scale, we show significant (p<0.05) overlap in genes identified for each aminoglycoside evaluated. Among the genomic segments identified, we confirmed increased resistance associated with an increased copy number of several genomic regions, including the ORF of PA5471, recently implicated in MexXY efflux pump related aminoglycoside resistance, PA4943-PA4946 (encoding a probable GTP-binding protein, a predicted host factor I protein, a delta 2-isopentenylpyrophosphate transferase, and DNA mismatch repair protein mutL), PA0960-PA0963 (encoding hypothetical proteins, a probable cold shock protein, a probable DNA-binding stress protein, and aspartyl-tRNA synthetase), a segment of PA4967 (encoding a topoisomerase IV subunit B), as well as a chimeric clone containing two inserts including the ORFs PA0547 and PA2326 (encoding a probable transcriptional regulator and a probable hypothetical protein, respectively).
Conclusions: The studies reported here demonstrate the application of new a genomic method, SCALEs, which can be used to improve understanding of the evolution of antibiotic resistance in P. aeruginosa. In our demonstration studies, we identified a significant number of genomic regions that increased resistance to multiple aminoglycosides. We identified genetic regions that include open reading frames that encode for products from many functional categories, including genes related to O-antigen synthesis, DNA repair, and transcriptional and translational processes.
Conflict of interest statement
Figures
Similar articles
-
Genomics of Aminoglycoside Resistance in Pseudomonas aeruginosa Bloodstream Infections at a United States Academic Hospital.Microbiol Spectr. 2023 Jun 15;11(3):e0508722. doi: 10.1128/spectrum.05087-22. Epub 2023 May 16. Microbiol Spectr. 2023. PMID: 37191517 Free PMC article.
-
Aminoglycoside resistance in Pseudomonas aeruginosa: the contribution of the MexXY-OprM efflux pump varies between isolates.J Med Microbiol. 2022 Jun;71(6). doi: 10.1099/jmm.0.001551. J Med Microbiol. 2022. PMID: 35708991
-
In vitro susceptibility of gentamicin-resistant Enterobacteriaceae and Pseudomonas aeruginosa to aminoglycoside antibiotics.Bol Asoc Med P R. 1983 Jul;75(7):305-7. Bol Asoc Med P R. 1983. PMID: 6442865 No abstract available.
-
Aminoglycosides plus beta-lactams against gram-negative organisms. Evaluation of in vitro synergy and chemical interactions.Am J Med. 1986 Jun 30;80(6B):126-37. doi: 10.1016/0002-9343(86)90490-0. Am J Med. 1986. PMID: 3088998 Review.
-
Antimicrobial agents--Part II. The aminoglycosides: streptomycin, kanamycin, gentamicin, tobramycin, amikacin, neomycin.Mayo Clin Proc. 1977 Nov;52(11):675-9. Mayo Clin Proc. 1977. PMID: 336988 Review.
Cited by
-
Intraspecific variation in antibiotic resistance potential within E. coli.Microbiol Spectr. 2024 Jun 4;12(6):e0316223. doi: 10.1128/spectrum.03162-23. Epub 2024 Apr 25. Microbiol Spectr. 2024. PMID: 38661581 Free PMC article.
-
Molecular Evolution of Extensively Drug-Resistant (XDR) Pseudomonas aeruginosa Strains From Patients and Hospital Environment in a Prolonged Outbreak.Front Microbiol. 2019 Aug 8;10:1742. doi: 10.3389/fmicb.2019.01742. eCollection 2019. Front Microbiol. 2019. PMID: 31440214 Free PMC article.
-
The Pseudomonas aeruginosa Resistome: Permanent and Transient Antibiotic Resistance, an Overview.Methods Mol Biol. 2024;2721:85-102. doi: 10.1007/978-1-0716-3473-8_7. Methods Mol Biol. 2024. PMID: 37819517
-
Genome-wide mapping of furfural tolerance genes in Escherichia coli.PLoS One. 2014 Jan 28;9(1):e87540. doi: 10.1371/journal.pone.0087540. eCollection 2014. PLoS One. 2014. PMID: 24489935 Free PMC article.
-
Recruitment of genes and enzymes conferring resistance to the nonnatural toxin bromoacetate.Proc Natl Acad Sci U S A. 2010 Oct 19;107(42):17968-73. doi: 10.1073/pnas.1007559107. Epub 2010 Oct 4. Proc Natl Acad Sci U S A. 2010. PMID: 20921376 Free PMC article.
References
-
- Lynch MD, Warnecke T, Gill RT. SCALEs: multiscale analysis of library enrichment. Nat Methods. 2007;4:87–93. - PubMed
-
- Leeb M. Antibiotics: a shot in the arm. Nature. 2004;431:892–893. - PubMed
-
- Wenzel RP. The antibiotic pipeline–challenges, costs, and values. N Engl J Med. 2004;351:523–526. - PubMed
-
- Courvalin P. Combinatorial approach of bacteria to antibiotic resistance. Res Microiology. 1999;150:1–7. - PubMed
-
- Baquero F. Low-level antibacterial resistance: a gateway to clinical resistance. Drug Resistance Updates. 2001;4:93–105. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

