A generic algorithm for layout of biological networks
- PMID: 19909528
- PMCID: PMC2785797
- DOI: 10.1186/1471-2105-10-375
A generic algorithm for layout of biological networks
Abstract
Background: Biological networks are widely used to represent processes in biological systems and to capture interactions and dependencies between biological entities. Their size and complexity is steadily increasing due to the ongoing growth of knowledge in the life sciences. To aid understanding of biological networks several algorithms for laying out and graphically representing networks and network analysis results have been developed. However, current algorithms are specialized to particular layout styles and therefore different algorithms are required for each kind of network and/or style of layout. This increases implementation effort and means that new algorithms must be developed for new layout styles. Furthermore, additional effort is necessary to compose different layout conventions in the same diagram. Also the user cannot usually customize the placement of nodes to tailor the layout to their particular need or task and there is little support for interactive network exploration.
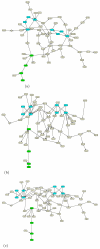
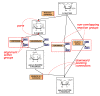
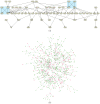
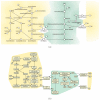
Results: We present a novel algorithm to visualize different biological networks and network analysis results in meaningful ways depending on network types and analysis outcome. Our method is based on constrained graph layout and we demonstrate how it can handle the drawing conventions used in biological networks.
Conclusion: The presented algorithm offers the ability to produce many of the fundamental popular drawing styles while allowing the exibility of constraints to further tailor these layouts.
Figures


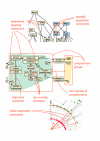
References
-
- Ehrenberg M, Elf J, Aurell E, Sandberg R, Tegnér J. Systems Biology Is Taking Off. Genome Research. 2003;13:2377–2380. - PubMed
-
- Kitano H. Systems Biology: A Brief Overview. Science. 2002;295(5560):1662–1664. - PubMed
-
- Basalaj W, Eilbeck K. In: Proc. International Symposium on Graph Drawing (GD'99), LNCS. Kratochvil J, editor. Vol. 1731. Stirin Castle: Springer; 1999. Straight-line drawings of protein interactions; pp. 259–266.
-
- Becker MY, Rojas I. A graph layout algorithm for drawing metabolic pathways. Bioinformatics. 2001;17(5):461–467. - PubMed
-
- Genc B, Dogrusöoz U. In: Proc. International Symposium on Graph Drawing (GD'03), LNCS. Liotta G, editor. Vol. 2912. Perugia: Springer; 2003. A Constrained, Force-Directed Layout Algorithm for Biological Pathways; pp. 314–319.
MeSH terms
LinkOut - more resources
Full Text Sources

