Viral genomic single-stranded RNA directs the pathway toward a T=3 capsid
- PMID: 19913556
- PMCID: PMC4785722
- DOI: 10.1016/j.jmb.2009.11.018
Viral genomic single-stranded RNA directs the pathway toward a T=3 capsid
Abstract
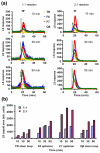
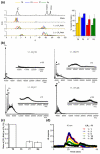
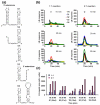
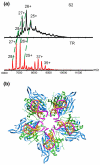
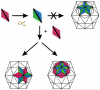
The molecular mechanisms controlling genome packaging by single-stranded RNA viruses are still largely unknown. It is necessary in most cases for the protein to adopt different conformations at different positions on the capsid lattice in order to form a viral capsid from multiple copies of a single protein. We showed previously that such quasi-equivalent conformers of RNA bacteriophage MS2 coat protein dimers (CP(2)) can be switched by sequence-specific interaction with a short RNA stem-loop (TR) that occurs only once in the wild-type phage genome. In principle, multiple switching events are required to generate the phage T=3 capsid. We have therefore investigated the sequence dependency of this event using two RNA aptamer sequences selected to bind the phage coat protein and an analogous packaging signal from phage Qbeta known to be discriminated against by MS2 coat protein both in vivo and in vitro. All three non-cognate stem-loops support T=3 shell formation, but none shows the kinetic-trapping effect seen when TR is mixed with equimolar CP(2). We show that this reflects the fact that they are poor ligands compared with TR, failing to saturate the coat protein under the assay conditions, ensuring that sufficient amounts of both types of dimer required for efficient assembly are present in these reactions. Increasing the non-cognate RNA concentration restores the kinetic trap, confirming this interpretation. We have also assessed the effects of extending the TR stem-loop at the 5' or 3' end with short genomic sequences. These longer RNAs all show evidence of the kinetic trap, reflecting the fact that they all contain the TR sequence and are more efficient at promoting capsid formation than TR. Mass spectrometry has shown that at least two pathways toward the T=3 shell occur in TR-induced assembly reactions: one via formation of a 3-fold axis and another that creates an extended 5-fold complex. The longer genomic RNAs suppress the 5-fold pathway, presumably as a consequence of steric clashes between multiply bound RNAs. Reversing the orientation of the extension sequences with respect to the TR stem-loop produces RNAs that are poor assembly initiators. The data support the idea that RNA-induced protein conformer switching occurs throughout assembly of the T=3 shell and show that both positional and sequence-specific effects outside the TR stem-loop can have significant impacts on the precise assembly pathway followed.
Copyright 2009 Elsevier Ltd. All rights reserved.
Figures

References
-
- Harrison SC, Olson AJ, Schutt CE, Winkler FK, Bricogne G. Tomato bushy stunt virus at 2.9 Å resolution. Nature. 1978;276:368–373. - PubMed
-
- Sorger PK, Stockley PG, Harrison SC. Structure and assembly of turnip crinkle virus: II. Mechanism of reassembly in vitro. J. Mol. Biol. 1986;191:639–658. - PubMed
-
- Caspar DL, Klug A. Physical principles in the construction of regular viruses. Cold Spring Harbor Symp. Quant. Biol. 1962;27:1–24. - PubMed
-
- Golmohammadi R, Valegard K, Fridborg K, Liljas L. The refined structure of bacteriophage MS2 at 2.8 Å resolution. J. Mol. Biol. 1993;234:620–639. - PubMed
-
- Valegard K, Liljas L, Fridborg K, Unge T. The three-dimensional structure of the bacterial virus MS2. Nature. 1990;345:36–41. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

