Beyond ribosome rescue: tmRNA and co-translational processes
- PMID: 19914241
- PMCID: PMC2794900
- DOI: 10.1016/j.febslet.2009.11.023
Beyond ribosome rescue: tmRNA and co-translational processes
Abstract
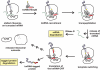
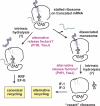
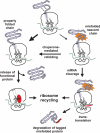
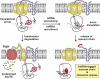
tmRNA is a unique bi-functional RNA that acts as both a tRNA and an mRNA to enter stalled ribosomes and direct the addition of a peptide tag to the C terminus of nascent polypeptides. Despite a reasonably clear understanding of tmRNA activity, the reason for its absolute conservation throughout the eubacteria is unknown. Although tmRNA plays many physiological roles in different bacterial systems, recent studies suggest a general role for trans-translation in monitoring protein folding and perhaps other co-translational processes. This review will focus on these new hypotheses and the data that support them.
Figures
References
-
- Barends S, Wower J, Kraal B. Kinetic parameters for tmRNA binding to alanyl-tRNA synthetase and elongation factor Tu from Escherichia coli. Biochemistry. 2000;39:2652–2658. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

