Algorithms for effective querying of compound graph-based pathway databases
- PMID: 19917102
- PMCID: PMC2784781
- DOI: 10.1186/1471-2105-10-376
Algorithms for effective querying of compound graph-based pathway databases
Abstract
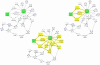
Background: Graph-based pathway ontologies and databases are widely used to represent data about cellular processes. This representation makes it possible to programmatically integrate cellular networks and to investigate them using the well-understood concepts of graph theory in order to predict their structural and dynamic properties. An extension of this graph representation, namely hierarchically structured or compound graphs, in which a member of a biological network may recursively contain a sub-network of a somehow logically similar group of biological objects, provides many additional benefits for analysis of biological pathways, including reduction of complexity by decomposition into distinct components or modules. In this regard, it is essential to effectively query such integrated large compound networks to extract the sub-networks of interest with the help of efficient algorithms and software tools.
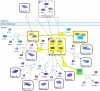
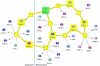
Results: Towards this goal, we developed a querying framework, along with a number of graph-theoretic algorithms from simple neighborhood queries to shortest paths to feedback loops, that is applicable to all sorts of graph-based pathway databases, from PPIs (protein-protein interactions) to metabolic and signaling pathways. The framework is unique in that it can account for compound or nested structures and ubiquitous entities present in the pathway data. In addition, the queries may be related to each other through "AND" and "OR" operators, and can be recursively organized into a tree, in which the result of one query might be a source and/or target for another, to form more complex queries. The algorithms were implemented within the querying component of a new version of the software tool PATIKAweb (Pathway Analysis Tool for Integration and Knowledge Acquisition) and have proven useful for answering a number of biologically significant questions for large graph-based pathway databases.
Conclusion: The PATIKA Project Web site is http://www.patika.org. PATIKAweb version 2.1 is available at http://web.patika.org.
Figures






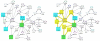
Similar articles
-
PATIKAweb: a Web interface for analyzing biological pathways through advanced querying and visualization.Bioinformatics. 2006 Feb 1;22(3):374-5. doi: 10.1093/bioinformatics/bti776. Epub 2005 Nov 15. Bioinformatics. 2006. PMID: 16287939
-
Integrated querying and version control of context-specific biological networks.Database (Oxford). 2020 Jan 1;2020:baaa018. doi: 10.1093/database/baaa018. Database (Oxford). 2020. PMID: 32294194 Free PMC article.
-
A query language for biological networks.Bioinformatics. 2005 Sep 1;21 Suppl 2:ii33-9. doi: 10.1093/bioinformatics/bti1105. Bioinformatics. 2005. PMID: 16204121
-
Graph databases in systems biology: a systematic review.Brief Bioinform. 2024 Sep 23;25(6):bbae561. doi: 10.1093/bib/bbae561. Brief Bioinform. 2024. PMID: 39565895 Free PMC article.
-
Graph theoretic modeling of large-scale semantic networks.J Biomed Inform. 2006 Aug;39(4):451-64. doi: 10.1016/j.jbi.2005.10.007. Epub 2005 Dec 15. J Biomed Inform. 2006. PMID: 16442849 Review.
Cited by
-
SyBLaRS: A web service for laying out, rendering and mining biological maps in SBGN, SBML and more.PLoS Comput Biol. 2022 Nov 14;18(11):e1010635. doi: 10.1371/journal.pcbi.1010635. eCollection 2022 Nov. PLoS Comput Biol. 2022. PMID: 36374853 Free PMC article.
-
STON: exploring biological pathways using the SBGN standard and graph databases.BMC Bioinformatics. 2016 Dec 5;17(1):494. doi: 10.1186/s12859-016-1394-x. BMC Bioinformatics. 2016. PMID: 27919219 Free PMC article.
-
Pathway Analysis with Signaling Hypergraphs.IEEE/ACM Trans Comput Biol Bioinform. 2017 Sep-Oct;14(5):1042-1055. doi: 10.1109/TCBB.2015.2459681. IEEE/ACM Trans Comput Biol Bioinform. 2017. PMID: 28991726 Free PMC article.
-
Integrating biological pathways and genomic profiles with ChiBE 2.BMC Genomics. 2014 Aug 3;15(1):642. doi: 10.1186/1471-2164-15-642. BMC Genomics. 2014. PMID: 25086704 Free PMC article.
-
Human protein reference database and human proteinpedia as discovery resources for molecular biotechnology.Mol Biotechnol. 2011 May;48(1):87-95. doi: 10.1007/s12033-010-9336-8. Mol Biotechnol. 2011. PMID: 20927658
References
-
- Matthews L, Gopinath G, Gillespie M, Caudy M, Croft D, de Bono B, Garapati P, Hemish J, Hermjakob H, Jassal B, Kanapin A, Lewis S, Mahajan S, May B, Schmidt E, Vastrik I, Wu G, Birney E, Stein L, D'Eustachio P. Reactome knowledgebase of human biological pathways and processes. Nucleic Acids Res. 2009. pp. D619–22. - DOI - PMC - PubMed
-
- Caspi R, Foerster H, Fulcher CA, Kaipa P, Krummenacker M, Latendresse M, Paley S, Rhee SY, Shearer AG, Tissier C, Walk TC, Zhang P, Karp PD. The MetaCyc Database of metabolic pathways and enzymes and the BioCyc collection of Pathway/Genome Databases. Nucleic Acids Res. 2008. pp. D623–31. - PMC - PubMed
-
- Bondy JA, Murty USR. Graph Theory with Applications. Great Britain: The McMillan Press Ltd; 1976.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

