E6 proteins from diverse papillomaviruses self-associate both in vitro and in vivo
- PMID: 19917295
- PMCID: PMC3900769
- DOI: 10.1016/j.jmb.2009.11.022
E6 proteins from diverse papillomaviruses self-associate both in vitro and in vivo
Erratum in
-
Corrigendum to "E6 Proteins from Diverse Papillomaviruses Self-Associate Both In Vitro and In Vivo" [J. Mol. Biol. 396(1) (2010) 90-104].J Mol Biol. 2026 Mar 1;438(5):169637. doi: 10.1016/j.jmb.2026.169637. Epub 2026 Jan 20. J Mol Biol. 2026. PMID: 41564712 No abstract available.
Abstract
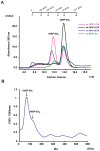
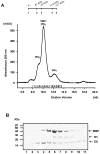
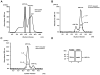
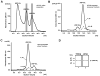
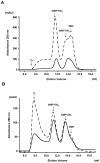
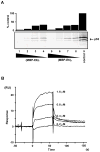
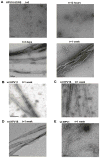
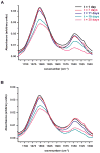
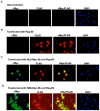
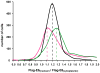
Papillomavirus E6 oncoproteins bind and often provoke the degradation of many cellular proteins important for the control of cell proliferation and/or cell death. Structural studies on E6 proteins have long been hindered by the difficulties of obtaining highly concentrated samples of recombinant E6. Here, we show that recombinant E6 proteins from eight human papillomavirus strains and one bovine papillomavirus strain exist as oligomeric and multimeric species. These species were characterized using a variety of biochemical and biophysical techniques, including analytical gel filtration, activity assays, surface plasmon resonance, electron microscopy and Fourier transform infrared spectroscopy. The characterization of E6 oligomers is facilitated by the fusion to the maltose binding protein, which slows the formation of higher-order multimeric species. The proportion of each oligomeric form varies depending on the viral strain considered. Oligomers appear to consist of folded units, which, in the case of high-risk mucosal human papillomavirus E6, retain binding to the ubiquitin ligase E6-associated protein and the capacity to degrade the proapoptotic protein p53. In addition to the small-size oligomers, E6 proteins spontaneously assemble into large organized multimeric structures, a process that is accompanied by a significant increase in the beta-sheet secondary structure content. Finally, co-localisation experiments using E6 equipped with different tags further demonstrate the occurrence of E6 self-association in eukaryotic cells. The ensemble of these data suggests that self-association is a general property of E6 proteins that occurs both in vitro and in vivo and might therefore be functionally relevant.
Copyright 2009 Elsevier Ltd. All rights reserved.
Figures
References
-
- zur Hausen H. Papillomaviruses in human cancers. Proc Assoc Am Physicians. 1999;111:581–587. - PubMed
-
- Bosch FX, Manos MM, Munoz N, Sherman M, Jansen AM, Peto J, Schiffman MH, Moreno V, Kurman R, Shah KV. Prevalence of human papillomavirus in cervical cancer: a worldwide perspective. International biological study on cervical cancer (IBSCC) Study Group. J Natl Cancer Inst. 1995;87:796–802. - PubMed
-
- Pfister H. Chapter 8: Human papillomavirus and skin cancer. J Natl Cancer Inst Monogr. 2003;31:52–56. - PubMed
-
- Lambert PF, Baker CC, Howley PM. The genetics of bovine papillomavirus type 1. Annu Rev Genet. 1988;22:235–58. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

