Fitness costs and stability of a high-level ciprofloxacin resistance phenotype in Salmonella enterica serotype enteritidis: reduced infectivity associated with decreased expression of Salmonella pathogenicity island 1 genes
- PMID: 19917752
- PMCID: PMC2798541
- DOI: 10.1128/AAC.00801-09
Fitness costs and stability of a high-level ciprofloxacin resistance phenotype in Salmonella enterica serotype enteritidis: reduced infectivity associated with decreased expression of Salmonella pathogenicity island 1 genes
Abstract
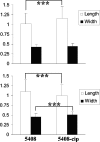
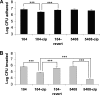
The fitness costs associated with high-level fluoroquinolone resistance were examined for phenotypically and genotypically characterized ciprofloxacin-resistant Salmonella enterica serotype Enteritidis mutants (104-cip and 5408-cip; MIC, >32 microg/ml). The stability of the fluoroquinolone resistance phenotype in both mutants was investigated to assess whether clones with better fitness could emerge in the absence of antibiotic selective pressure. Mutants 104-cip and 5408-cip displayed altered morphology on agar and by electron microscopy, reduced growth rates, motility and invasiveness in Caco-2 cells, and increased sensitivity to environmental stresses. Microarray data revealed decreased expression of virulence and motility genes in both mutants. Two clones, 104-revert and 1A-revertC2, with ciprofloxacin MICs of 3 and 2 microg/ml, respectively, were recovered from separate lineages of 104-cip after 20 and 70 passages, respectively, on antibiotic-free agar. All fitness costs, except motility, were reversed in 104-revert. Potential mechanisms associated with reversal of the resistance phenotype were examined. Compared to 104-cip, both 104-revert and 1A-revertC2 showed decreased expression of acrB and soxS but still overexpressed marA. Both acquired additional mutations in SoxR and ParC, and 1A-revertC2 acquired two mutations in MarA. The altered porin and lipopolysaccharide (LPS) profiles observed in 104-cip were reversed. In contrast, 5408-cip showed no reversal in fitness costs and maintained its high-level ciprofloxacin resistance for 200 passages on antibiotic-free agar. In conclusion, high-level ciprofloxacin resistance in S. Enteritidis is associated with fitness costs. In the absence of antibiotic selection pressure, isolates may acquire mutations enabling reversion to an intermediate-level ciprofloxacin resistance phenotype associated with less significant fitness costs.
Figures

References
-
- Alonso, A., G. Morales, R. Escalante, E. Campanario, L. Sastre, and J. L. Martinez. 2004. Overexpression of the multidrug efflux pump SmeDEF impairs Stenotrophomonas maltophilia physiology. J. Antimicrob. Chemother. 53:432-434. - PubMed
-
- Andersson, D. I. 2006. The biological cost of mutational antibiotic resistance: any practical conclusions? Curr. Opin. Microbiol. 9:461-465. - PubMed
-
- Carrique-Mas, J. J., C. Papadopoulou, S. J. Evans, A. Wales, C. J. Teale, and R. H. Davies. 2008. Trends in phage types and antimicrobial resistance of Salmonella enterica serovar Enteritidis isolated from animals in Great Britain from 1990 to 2005. Vet. Rec. 162:541-546. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

