Evaluating the performance of new approaches to spot quantification and differential expression in 2-dimensional gel electrophoresis studies
- PMID: 19919108
- PMCID: PMC2802214
- DOI: 10.1021/pr9005603
Evaluating the performance of new approaches to spot quantification and differential expression in 2-dimensional gel electrophoresis studies
Abstract
Spot detection and quantification for 2-DE are challenging and important tasks to fully extract the proteomic information from these data. Traditional analytical methods have significant weaknesses, including spot mismatching and missing data, which require time-consuming manual editing to correct, dramatically decreasing throughput and compromising the objectivity and reproducibility of the analysis. To address this issue, we developed Pinnacle, a novel, quick, automatic, noncommercial method that borrows strength across gels in spot detection and has been shown to yield more precise spot quantifications than traditional methods. New commercial software, notably SameSpots, has also recently been developed as an improvement over traditional workflows. In this paper, we briefly describe Pinnacle and compare its performance to SameSpots in spot detection, spot quantification precision, and differential expression. Our analysis is performed in a rigorous fashion that, unlike other comparisons in the literature, summarizes performance across all spots detected on the gels, and we manually optimize SameSpots results while simply running Pinnacle with standard settings and no manual editing. While both methods showed marked improvement over a commercially available traditional method PG240, Pinnacle consistently yielded spot quantifications with greater validity and reliability, avoided spot delineation problems, and detected more differentially expressed proteins than SameSpots, and represents a significant noncommercial alternative for 2-DE processing.
Conflict of interest statement
The authors do not have any financial or commercial conflicts of interest regarding this paper.
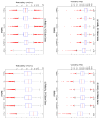
Figures

Similar articles
-
Detection and Quantification of Protein Spots by Pinnacle.Methods Mol Biol. 2016;1384:185-201. doi: 10.1007/978-1-4939-3255-9_11. Methods Mol Biol. 2016. PMID: 26611416 Free PMC article.
-
In the eye of the beholder: does the master see the SameSpots as the novice?J Proteome Res. 2010 Mar 5;9(3):1522-32. doi: 10.1021/pr9010298. J Proteome Res. 2010. PMID: 20108985
-
Pinnacle: a fast, automatic and accurate method for detecting and quantifying protein spots in 2-dimensional gel electrophoresis data.Bioinformatics. 2008 Feb 15;24(4):529-36. doi: 10.1093/bioinformatics/btm590. Epub 2008 Jan 14. Bioinformatics. 2008. PMID: 18194961 Free PMC article.
-
Comparison of two academic software packages for analyzing two-dimensional gel images.J Bioinform Comput Biol. 2011 Dec;9(6):775-94. doi: 10.1142/s0219720011005665. J Bioinform Comput Biol. 2011. PMID: 22084013 Review.
-
Geometrical distortions in two-dimensional gels: applicable correction methods.J Chromatogr B Analyt Technol Biomed Life Sci. 2005 Feb 5;815(1-2):25-37. doi: 10.1016/j.jchromb.2004.07.037. J Chromatogr B Analyt Technol Biomed Life Sci. 2005. PMID: 15652796 Review.
Cited by
-
Statistical Contributions to Bioinformatics: Design, Modeling, Structure Learning, and Integration.Stat Modelling. 2017;17(4-5):245-289. doi: 10.1177/1471082X17698255. Epub 2017 Jun 15. Stat Modelling. 2017. PMID: 29129969 Free PMC article.
-
Emmental Cheese Environment Enhances Propionibacterium freudenreichii Stress Tolerance.PLoS One. 2015 Aug 14;10(8):e0135780. doi: 10.1371/journal.pone.0135780. eCollection 2015. PLoS One. 2015. PMID: 26275229 Free PMC article.
-
Hyperconcentrated Sweet Whey, a New Culture Medium That Enhances Propionibacterium freudenreichii Stress Tolerance.Appl Environ Microbiol. 2016 Jul 15;82(15):4641-4651. doi: 10.1128/AEM.00748-16. Print 2016 Aug 1. Appl Environ Microbiol. 2016. PMID: 27235433 Free PMC article.
-
Detection and Quantification of Protein Spots by Pinnacle.Methods Mol Biol. 2016;1384:185-201. doi: 10.1007/978-1-4939-3255-9_11. Methods Mol Biol. 2016. PMID: 26611416 Free PMC article.
-
Two-Dimensional Gel Electrophoresis Image Analysis.Methods Mol Biol. 2021;2361:3-13. doi: 10.1007/978-1-0716-1641-3_1. Methods Mol Biol. 2021. PMID: 34236652
References
-
- Cutler P, Heald G, White IR, Ruan J. A novel approach to spot detection for two-dimensional gel electrophoresis images using pixel value collection. Proteomics. 2003;3:392–401. - PubMed
-
- Clark BN, Gutstein HB. The myth of automated, high-throughput two-dimensional gel electrophoresis. Proteomics. 2008;8:1197–1203. - PubMed
-
- Karp NA, Feret R, Rubtsov DV, Lilley KS. Comparison of DIGE and post-stained gel electrophoresis with both traditional and SameSpots analysis for quantitative proteomics. Proteomics. 2008;8:948–960. - PubMed
-
- Nishihara JC, Champion KM. Quantitative evaluation of proteins in one- and two-dimensional polyacrylamide gels using a fluorescent stain. Electrophoresis. 2002;23:2203–2215. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

