The Arabidopsis B-box zinc finger family
- PMID: 19920209
- PMCID: PMC2798317
- DOI: 10.1105/tpc.109.069088
The Arabidopsis B-box zinc finger family
Figures


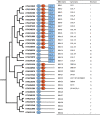
References
-
- Chang, C.S., Li, Y.H., Chen, L.-T., Chen, W.-C., Hsieh, W.-P., Shin, J., Jane, W.-N., Chou, S.-J., Choi, G., Hu, J.-M., Somerville, S., and Wu, S.-H. (2008). LZF1, a HY5-regulated transcriptional factor, functions in Arabidopsis de-etiolation. Plant J. 54 205–219. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

