A family of microRNAs encoded by myosin genes governs myosin expression and muscle performance
- PMID: 19922871
- PMCID: PMC2796371
- DOI: 10.1016/j.devcel.2009.10.013
A family of microRNAs encoded by myosin genes governs myosin expression and muscle performance
Abstract
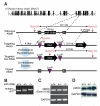
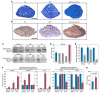
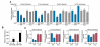
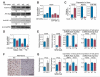
Myosin is the primary regulator of muscle strength and contractility. Here we show that three myosin genes, Myh6, Myh7, and Myh7b, encode related intronic microRNAs (miRNAs), which, in turn, control muscle myosin content, myofiber identity, and muscle performance. Within the adult heart, the Myh6 gene, encoding a fast myosin, coexpresses miR-208a, which regulates the expression of two slow myosins and their intronic miRNAs, Myh7/miR-208b and Myh7b/miR-499, respectively. miR-208b and miR-499 play redundant roles in the specification of muscle fiber identity by activating slow and repressing fast myofiber gene programs. The actions of these miRNAs are mediated in part by a collection of transcriptional repressors of slow myofiber genes. These findings reveal that myosin genes not only encode the major contractile proteins of muscle, but act more broadly to influence muscle function by encoding a network of intronic miRNAs that control muscle gene expression and performance.
Figures




References
-
- Adolph EA, Subramaniam A, Cserjesi P, Olson EN, Robbins J. Role of myocyte-specific enhancer-binding factor (MEF-2) in transcriptional regulation of the alpha-cardiac myosin heavy chain gene. J Biol Chem. 1993;268:5349–5352. - PubMed
-
- Azakie A, Fineman JR, He Y. Sp3 inhibits Sp1-mediated activation of the cardiac troponin T promoter and is downregulated during pathological cardiac hypertrophy in vivo. Am J Physiol Heart Circ Physiol. 2006;291:H600–611. - PubMed
-
- Baldwin KM, Haddad F. Effects of different activity and inactivity paradigms on myosin heavy chain gene expression in striated muscle. J Appl Physiol. 2001;90:345–357. - PubMed
-
- Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–297. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

