Systems-level dynamic analyses of fate change in murine embryonic stem cells
- PMID: 19924215
- PMCID: PMC3199216
- DOI: 10.1038/nature08575
Systems-level dynamic analyses of fate change in murine embryonic stem cells
Abstract
Molecular regulation of embryonic stem cell (ESC) fate involves a coordinated interaction between epigenetic, transcriptional and translational mechanisms. It is unclear how these different molecular regulatory mechanisms interact to regulate changes in stem cell fate. Here we present a dynamic systems-level study of cell fate change in murine ESCs following a well-defined perturbation. Global changes in histone acetylation, chromatin-bound RNA polymerase II, messenger RNA (mRNA), and nuclear protein levels were measured over 5 days after downregulation of Nanog, a key pluripotency regulator. Our data demonstrate how a single genetic perturbation leads to progressive widespread changes in several molecular regulatory layers, and provide a dynamic view of information flow in the epigenome, transcriptome and proteome. We observe that a large proportion of changes in nuclear protein levels are not accompanied by concordant changes in the expression of corresponding mRNAs, indicating important roles for translational and post-translational regulation of ESC fate. Gene-ontology analysis across different molecular layers indicates that although chromatin reconfiguration is important for altering cell fate, it is preceded by transcription-factor-mediated regulatory events. The temporal order of gene expression alterations shows the order of the regulatory network reconfiguration and offers further insight into the gene regulatory network. Our studies extend the conventional systems biology approach to include many molecular species, regulatory layers and temporal series, and underscore the complexity of the multilayer regulatory mechanisms responsible for changes in protein expression that determine stem cell fate.
Figures


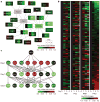

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

