Protein-precursor tRNA contact leads to sequence-specific recognition of 5' leaders by bacterial ribonuclease P
- PMID: 19932118
- PMCID: PMC2829246
- DOI: 10.1016/j.jmb.2009.11.039
Protein-precursor tRNA contact leads to sequence-specific recognition of 5' leaders by bacterial ribonuclease P
Abstract
Bacterial ribonuclease P (RNase P) catalyzes the cleavage of 5' leader sequences from precursor tRNAs (pre-tRNAs). Previously, all known substrate nucleotide specificities in this system are derived from RNA-RNA interactions with the RNase P RNA subunit. Here, we demonstrate that pre-tRNA binding affinities for Bacillus subtilis and Escherichia coli RNase P are enhanced by sequence-specific contacts between the fourth pre-tRNA nucleotide on the 5' side of the cleavage site (N(-4)) and the RNase P protein (P protein) subunit. B. subtilis RNase P has a higher affinity for pre-tRNA with adenosine at N(-4), and this binding preference is amplified at physiological divalent ion concentrations. Measurements of pre-tRNA-containing adenosine analogs at N(-4) indicate that specificity arises from a combination of hydrogen bonding to the N6 exocyclic amine of adenosine and steric exclusion of the N2 amine of guanosine. Mutagenesis of B. subtilis P protein indicates that F20 and Y34 contribute to selectivity at N(-4). The hydroxyl group of Y34 enhances selectivity, likely by forming a hydrogen bond with the N(-4) nucleotide. The sequence preference of E. coli RNase P is diminished, showing a weak preference for adenosine and cytosine at N(-4), consistent with the substitution of Leu for Y34 in the E. coli P protein. This is the first identification of a sequence-specific contact between P protein and pre-tRNA that contributes to molecular recognition of RNase P. Additionally, sequence analyses reveal that a greater-than-expected fraction of pre-tRNAs from both E. coli and B. subtilis contains a nucleotide at N(-4) that enhances RNase P affinity. This observation suggests that specificity at N(-4) contributes to substrate recognition in vivo. Furthermore, bioinformatic analyses suggest that sequence-specific contacts between the protein subunit and the leader sequences of pre-tRNAs may be common in bacterial RNase P and may lead to species-specific substrate recognition.
Copyright 2009 Elsevier Ltd. All rights reserved.
Figures

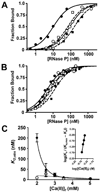



References
-
- Smith JK, Hsieh J, Fierke CA. Importance of RNA-protein interactions in bacterial ribonuclease P structure and catalysis. Biopolymers. 2007;87:329–338. - PubMed
-
- Kirsebom LA. RNase P RNA mediated cleavage: substrate recognition and catalysis. Biochimie. 2007;89:1183–1194. - PubMed
-
- Christian EL, Zahler NH, Kaye NM, Harris ME. Analysis of substrate recognition by the ribonucleoprotein endonuclease RNase P. Methods. 2002;28:307–322. - PubMed
-
- Beebe JA, Fierke CA. A kinetic mechanism for cleavage of precursor tRNA(Asp) catalyzed by the RNA component of Bacillus subtilis ribonuclease P. Biochemistry. 1994;33:10294–10304. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

