MouseIndelDB: a database integrating genomic indel polymorphisms that distinguish mouse strains
- PMID: 19933259
- PMCID: PMC2808983
- DOI: 10.1093/nar/gkp1046
MouseIndelDB: a database integrating genomic indel polymorphisms that distinguish mouse strains
Abstract
MouseIndelDB is an integrated database resource containing thousands of previously unreported mouse genomic indel (insertion and deletion) polymorphisms ranging from approximately 100 nt to 10 Kb in size. The database currently includes polymorphisms identified from our alignment of 26 million whole-genome shotgun sequence traces from four laboratory mouse strains mapped against the reference C57BL/6J genome using GMAP. They can be queried on a local level by chromosomal coordinates, nearby gene names or other genomic feature identifiers, or in bulk format using categories including mouse strain(s), class of polymorphism(s) and chromosome number. The results of such queries are presented either as a custom track on the UCSC mouse genome browser or in tabular format. We anticipate that the MouseIndelDB database will be widely useful for research in mammalian genetics, genomics, and evolutionary biology. Access to the MouseIndelDB database is freely available at: http://variation.osu.edu/.
Figures
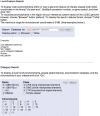

References
-
- Beck JA, Lloyd S, Hafezparast M, Lennon-Pierce M, Eppig JT, Festing MF, Fisher EM. Genealogies of mouse inbred strains. Nat. Genet. 2000;24:23–25. - PubMed
-
- Churchill GA, Airey DC, Allayee H, Angel JM, Attie AD, Beatty J, Beavis WD, Belknap JK, Bennett B, Berrettini W, et al. The Collaborative Cross, a community resource for the genetic analysis of complex traits. Nat. Genet. 2004;36:1133–1137. - PubMed
-
- Varki A, Altheide TK. Comparing the human and chimpanzee genomes: searching for needles in a haystack. Genome Res. 2005;15:1746–1758. - PubMed
-
- Wade CM, Daly MJ. Genetic variation in laboratory mice. Nat. Genet. 2005;37:1175–1180. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

