Stem cell signaling in Arabidopsis requires CRN to localize CLV2 to the plasma membrane
- PMID: 19933383
- PMCID: PMC2799354
- DOI: 10.1104/pp.109.149930
Stem cell signaling in Arabidopsis requires CRN to localize CLV2 to the plasma membrane
Abstract
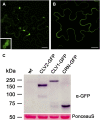
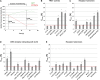
Stem cell number in shoot and floral meristems of Arabidopsis (Arabidopsis thaliana) is regulated by the CLAVATA3 (CLV3) signaling pathway. Perception of the CLV3 peptide requires the receptor kinase CLV1, the receptor-like protein CLV2, and the kinase CORYNE (CRN). Genetic analysis suggested that CLV2 and CRN act together and in parallel with CLV1. We studied the intracellular localization of receptor fusions with fluorescent protein tags and their capacities for interaction via efficiency of fluorescence resonance energy transfer. We found that CLV2 and CRN require each other for export from the endoplasmic reticulum and localization to the plasma membrane (PM). CRN readily forms homomers and interacts with CLV2 through the transmembrane domain and adjacent juxtamembrane sequences. CLV1 forms homomers independently of CLV2 and CRN at the PM. We propose that the CLV3 signal is perceived by a tetrameric CLV2/CRN complex and a CLV1 homodimer that localize to the PM and can interact via CRN.
Figures




References
-
- Albertazzi L, Arosio D, Marchetti L, Ricci F, Beltram F (2009) Quantitative FRET analysis with the EGFP-mCherry fluorescent protein pair. Photochem Photobiol 85 287–297 - PubMed
-
- Ali GS, Prasad KV, Day I, Reddy AS (2007) Ligand-dependent reduction in the membrane mobility of FLAGELLIN SENSITIVE2, an Arabidopsis receptor-like kinase. Plant Cell Physiol 48 1601–1611 - PubMed
-
- Boller T, Felix G (2009) A renaissance of elicitors: perception of microbe-associated molecular patterns and danger signals by pattern-recognition receptors. Annu Rev Plant Biol 60 379–406 - PubMed
-
- Brand U, Fletcher JC, Hobe M, Meyerowitz EM, Simon R (2000) Dependence of stem cell fate in Arabidopsis on a feedback loop regulated by CLV3 activity. Science 289 617–619 - PubMed
-
- Chan FK, Chun HJ, Zheng L, Siegel RM, Bui KL, Lenardo MJ (2000) A domain in TNF receptors that mediates ligand-independent receptor assembly and signaling. Science 288 2351–2354 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

