The nuclear component of a cytonuclear hybrid incompatibility in Mimulus maps to a cluster of pentatricopeptide repeat genes
- PMID: 19933877
- PMCID: PMC2828725
- DOI: 10.1534/genetics.109.108175
The nuclear component of a cytonuclear hybrid incompatibility in Mimulus maps to a cluster of pentatricopeptide repeat genes
Abstract
Characterizing the genetic and molecular basis of hybrid incompatibilities is a first step toward understanding their evolutionary origins. We fine mapped the nuclear restorer (Rf) of cytoplasm-dependent anther sterility in Mimulus hybrids by identifying and targeting regions of the Mimulus guttatus genome containing large numbers of candidate pentatricopeptide repeat genes (PPRs). The single Mendelian locus Rf was first isolated to a 1.3-cM region on linkage group 7 that spans the genome's largest cluster of PPRs, then split into two tightly linked loci (Rf1 and Rf2) by <10 recombination events in a large (N = 6153) fine-mapping population. Progeny testing of fertile recombinants demonstrated that a dominant M. guttatus allele at each Rf locus was sufficient to restore fertility. Each Rf locus spans a physical region containing numerous PPRs with high homology to each other, suggesting recent tandem duplication or transposition. Furthermore, these PPRs have higher homology to restorers in distantly related taxa (petunia and rice) than to PPRs elsewhere in the Mimulus genome. These results suggest that the cytoplasmic male sterility (CMS)-PPR interaction is highly conserved across flowering plants. In addition, given our theoretical understanding of cytonuclear coevolution, the finding that hybrid CMS results from interactions between a chimeric mitochondrial transcript that is modified by Rf loci identified as PPRs is consistent with a history of selfish mitochondrial evolution and compensatory nuclear coevolution within M. guttatus.
Figures
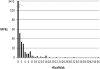

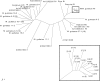
References
-
- Abascal, F., R. Zardoya and D. Posada, 2005. ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21 2104–2105. - PubMed
-
- Altschul, S. F., W. Gish, W. Miller, E. W. Myers and D. J. Lipman, 1990. Basic local alignment search tool. J. Mol. Biol. 215 403–410. - PubMed
-
- Andres, C., C. Lurin and I. D. Small, 2007. The multifarious roles of PPR proteins in plant mitochondrial gene expression. Physiol. Plant. 129 14–22.
-
- Ashman, T. L., 1994. Reproductive allocation in hermaphrodite and female plants of Sidalcea oregana spp. spicata (Malvaceae) using 4 currencies. Am. J. Bot. 81 433–438.
-
- Atlan, A., P. H. Gouyon, T. Fournial, D. Pomente and D. Couvet, 1992. Sex allocation in an hermaphroditic plant: the case of gynodioecy in Thymus vulgaris L. J. Evol. Biol. 5 189–203.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

