Detailed analysis of a contiguous 22-Mb region of the maize genome
- PMID: 19936048
- PMCID: PMC2773423
- DOI: 10.1371/journal.pgen.1000728
Detailed analysis of a contiguous 22-Mb region of the maize genome
Abstract
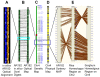
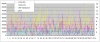
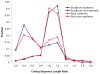
Most of our understanding of plant genome structure and evolution has come from the careful annotation of small (e.g., 100 kb) sequenced genomic regions or from automated annotation of complete genome sequences. Here, we sequenced and carefully annotated a contiguous 22 Mb region of maize chromosome 4 using an improved pseudomolecule for annotation. The sequence segment was comprehensively ordered, oriented, and confirmed using the maize optical map. Nearly 84% of the sequence is composed of transposable elements (TEs) that are mostly nested within each other, of which most families are low-copy. We identified 544 gene models using multiple levels of evidence, as well as five miRNA genes. Gene fragments, many captured by TEs, are prevalent within this region. Elimination of gene redundancy from a tetraploid maize ancestor that originated a few million years ago is responsible in this region for most disruptions of synteny with sorghum and rice. Consistent with other sub-genomic analyses in maize, small RNA mapping showed that many small RNAs match TEs and that most TEs match small RNAs. These results, performed on approximately 1% of the maize genome, demonstrate the feasibility of refining the B73 RefGen_v1 genome assembly by incorporating optical map, high-resolution genetic map, and comparative genomic data sets. Such improvements, along with those of gene and repeat annotation, will serve to promote future functional genomic and phylogenomic research in maize and other grasses.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
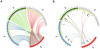
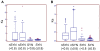
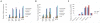
Similar articles
-
Diverged copies of the seed regulatory Opaque-2 gene by a segmental duplication in the progenitor genome of rice, sorghum, and maize.Mol Plant. 2008 Sep;1(5):760-9. doi: 10.1093/mp/ssn038. Epub 2008 Aug 1. Mol Plant. 2008. PMID: 19825579
-
The physical and genetic framework of the maize B73 genome.PLoS Genet. 2009 Nov;5(11):e1000715. doi: 10.1371/journal.pgen.1000715. Epub 2009 Nov 20. PLoS Genet. 2009. PMID: 19936061 Free PMC article.
-
Sequencing, mapping, and analysis of 27,455 maize full-length cDNAs.PLoS Genet. 2009 Nov;5(11):e1000740. doi: 10.1371/journal.pgen.1000740. Epub 2009 Nov 20. PLoS Genet. 2009. PMID: 19936069 Free PMC article.
-
Maize as a model for the evolution of plant nuclear genomes.Proc Natl Acad Sci U S A. 2000 Jun 20;97(13):7008-15. doi: 10.1073/pnas.97.13.7008. Proc Natl Acad Sci U S A. 2000. PMID: 10860964 Free PMC article. Review.
-
Tracking footprints of maize domestication and evidence for a massive selective sweep on chromosome 10.Proc Natl Acad Sci U S A. 2009 Jun 16;106 Suppl 1(Suppl 1):9979-86. doi: 10.1073/pnas.0901122106. Epub 2009 Jun 15. Proc Natl Acad Sci U S A. 2009. PMID: 19528660 Free PMC article. Review.
Cited by
-
Quadruplex-Forming Motif Inserted into 3'UTR of Ty1his3-AI Retrotransposon Inhibits Retrotransposition in Yeast.Biology (Basel). 2021 Apr 20;10(4):347. doi: 10.3390/biology10040347. Biology (Basel). 2021. PMID: 33924086 Free PMC article.
-
Landscape of next-generation sequencing technologies.Anal Chem. 2011 Jun 15;83(12):4327-41. doi: 10.1021/ac2010857. Epub 2011 May 25. Anal Chem. 2011. PMID: 21612267 Free PMC article. Review. No abstract available.
-
Variation and Inheritance of Small RNAs in Maize Inbreds and F1 Hybrids.Plant Physiol. 2020 Jan;182(1):318-331. doi: 10.1104/pp.19.00817. Epub 2019 Oct 1. Plant Physiol. 2020. PMID: 31575624 Free PMC article.
-
The barley Frost resistance-H2 locus.Funct Integr Genomics. 2014 Mar;14(1):85-100. doi: 10.1007/s10142-014-0360-9. Epub 2014 Jan 18. Funct Integr Genomics. 2014. PMID: 24442711
-
miRanalyzer: an update on the detection and analysis of microRNAs in high-throughput sequencing experiments.Nucleic Acids Res. 2011 Jul;39(Web Server issue):W132-8. doi: 10.1093/nar/gkr247. Epub 2011 Apr 22. Nucleic Acids Res. 2011. PMID: 21515631 Free PMC article.
References
-
- Duvick DN. Genetic contributions to yield gains of U.S. hybrid maize, 1930–1980. In: Fehr WR, editor. Genetic contributions to yield gains of five major crop plants. Madison, WI: Crop Science Soc of America Special Publ No 7; 1984.
-
- Bennetzen JL, Hake S. The maize handbook; Volumes 1 and II. New York: Springer; 2009.
-
- McClintock B. Mutable loci in maize. Carnegie Inst Wash Year Book. 1948;47:155–169. - PubMed
-
- SanMiguel P, Tikhonov A, Jin Y-K, Motchoulskaia N, Zakharov D, et al. Nested retrotransposons in the intergenic regions of the maize genome. Science. 1996;274:765–768. - PubMed
-
- SanMiguel P, Bennetzen JL. Evidence that a recent increase in maize genome size was caused by the massive amplification of intergene retrotransposons. Annals Bot. 1998;82:37–44.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

