Sequencing, mapping, and analysis of 27,455 maize full-length cDNAs
- PMID: 19936069
- PMCID: PMC2774520
- DOI: 10.1371/journal.pgen.1000740
Sequencing, mapping, and analysis of 27,455 maize full-length cDNAs
Abstract
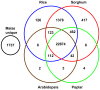
Full-length cDNA (FLcDNA) sequencing establishes the precise primary structure of individual gene transcripts. From two libraries representing 27 B73 tissues and abiotic stress treatments, 27,455 high-quality FLcDNAs were sequenced. The average transcript length was 1.44 kb including 218 bases and 321 bases of 5' and 3' UTR, respectively, with 8.6% of the FLcDNAs encoding predicted proteins of fewer than 100 amino acids. Approximately 94% of the FLcDNAs were stringently mapped to the maize genome. Although nearly two-thirds of this genome is composed of transposable elements (TEs), only 5.6% of the FLcDNAs contained TE sequences in coding or UTR regions. Approximately 7.2% of the FLcDNAs are putative transcription factors, suggesting that rare transcripts are well-enriched in our FLcDNA set. Protein similarity searching identified 1,737 maize transcripts not present in rice, sorghum, Arabidopsis, or poplar annotated genes. A strict FLcDNA assembly generated 24,467 non-redundant sequences, of which 88% have non-maize protein matches. The FLcDNAs were also assembled with 41,759 FLcDNAs in GenBank from other projects, where semi-strict parameters were used to identify 13,368 potentially unique non-redundant sequences from this project. The libraries, ESTs, and FLcDNA sequences produced from this project are publicly available. The annotated EST and FLcDNA assemblies are available through the maize FLcDNA web resource (www.maizecdna.org).
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

Similar articles
-
Analysis of 4,664 high-quality sequence-finished poplar full-length cDNA clones and their utility for the discovery of genes responding to insect feeding.BMC Genomics. 2008 Jan 29;9:57. doi: 10.1186/1471-2164-9-57. BMC Genomics. 2008. PMID: 18230180 Free PMC article.
-
A conifer genomics resource of 200,000 spruce (Picea spp.) ESTs and 6,464 high-quality, sequence-finished full-length cDNAs for Sitka spruce (Picea sitchensis).BMC Genomics. 2008 Oct 14;9:484. doi: 10.1186/1471-2164-9-484. BMC Genomics. 2008. PMID: 18854048 Free PMC article.
-
Computational finishing of large sequence contigs reveals interspersed nested repeats and gene islands in the rf1-associated region of maize.Plant Physiol. 2009 Oct;151(2):483-95. doi: 10.1104/pp.109.143370. Epub 2009 Aug 12. Plant Physiol. 2009. PMID: 19675151 Free PMC article.
-
Progress in maize gene discovery: a project update.Funct Integr Genomics. 2003 Mar;3(1-2):25-32. doi: 10.1007/s10142-002-0078-y. Epub 2002 Oct 1. Funct Integr Genomics. 2003. PMID: 12590340 Review.
-
Assembling genomes using short-read sequencing technology.Genome Biol. 2010 Jan 28;11(1):202. doi: 10.1186/gb-2010-11-1-202. Genome Biol. 2010. PMID: 20128932 Free PMC article. Review.
Cited by
-
RAMOSA1 ENHANCER LOCUS2-Mediated Transcriptional Repression Regulates Vegetative and Reproductive Architecture.Plant Physiol. 2019 Jan;179(1):348-363. doi: 10.1104/pp.18.00913. Epub 2018 Oct 22. Plant Physiol. 2019. PMID: 30348817 Free PMC article.
-
10 reasons to be tantalized by the B73 maize genome.PLoS Genet. 2009 Nov;5(11):e1000723. doi: 10.1371/journal.pgen.1000723. Epub 2009 Nov 20. PLoS Genet. 2009. PMID: 19936060 Free PMC article. No abstract available.
-
An expression analysis of 57 transcription factors derived from ESTs of developing seeds in Maize (Zea mays).Plant Cell Rep. 2010 Jun;29(6):545-59. doi: 10.1007/s00299-010-0843-7. Epub 2010 Mar 25. Plant Cell Rep. 2010. PMID: 20336461
-
Establishing a System for Functional Characterization of Full-Length cDNAs of Camellia sinensis.Int J Mol Sci. 2019 Nov 25;20(23):5929. doi: 10.3390/ijms20235929. Int J Mol Sci. 2019. PMID: 31775391 Free PMC article.
-
A k-mer grammar analysis to uncover maize regulatory architecture.BMC Plant Biol. 2019 Mar 15;19(1):103. doi: 10.1186/s12870-019-1693-2. BMC Plant Biol. 2019. PMID: 30876396 Free PMC article.
References
-
- Buckler ES, Gaut BS, McMullen MD. Molecular and functional diversity of maize. Curr Opin Plant Biol. 2006;9:172–176. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

