The structure and NO binding properties of the nitrophorin-like heme-binding protein from Arabidopsis thaliana gene locus At1g79260.1
- PMID: 19938152
- PMCID: PMC2811769
- DOI: 10.1002/prot.22617
The structure and NO binding properties of the nitrophorin-like heme-binding protein from Arabidopsis thaliana gene locus At1g79260.1
Abstract
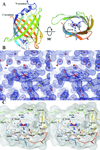
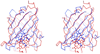
The protein from Arabidopsis thaliana gene locus At1g79260.1 is comprised of 166-residues and is of previously unknown function. Initial structural studies by the Center for Eukaryotic Structural Genomics (CESG) suggested that this protein might bind heme, and consequently, the crystal structures of apo and heme-bound forms were solved to near atomic resolution of 1.32 A and 1.36 A, respectively. The rate of hemin loss from the protein was measured to be 3.6 x 10(-5) s(-1), demonstrating that it binds heme specifically and with high affinity. The protein forms a compact 10-stranded beta-barrel that is structurally similar to the lipocalins and fatty acid binding proteins (FABPs). One group of lipocalins, the nitrophorins (NP), are heme proteins involved in nitric oxide (NO) transport and show both sequence and structural similarity to the protein from At1g79260.1 and two human homologues, all of which contain a proximal histidine capable of coordinating a heme iron. Rapid-mixing and laser photolysis techniques were used to determine the rate constants for carbon monoxide (CO) binding to the ferrous form of the protein (k'(CO) = 0.23 microM(-1) s(-1), k(CO) = 0.050 s(-1)) and NO binding to the ferric form (k'(NO) = 1.2 microM(-1) s(-1), k(NO) = 73 s(-1)). Based on both structural and functional similarity to the nitrophorins, we have named the protein nitrobindin and hypothesized that it plays a role in NO transport. However, one of the two human homologs of nitrobindin contains a THAP domain, implying a possible role in apoptosis. Proteins 2010. (c) 2009 Wiley-Liss, Inc.
Figures




References
-
- Gray HB, Winkler JR. Electron transfer in proteins. Annu Rev Biochem. 1996;65:537–561. - PubMed
-
- Ortiz de Montellano PR. Cytochrome P450: Structure, Mechanism, and Biochemistry. New York: Plenium press; 1995.
-
- Rodgers KR. Heme-based sensors in biological systems. Curr Opin Chem Biol. 1999;3(2):158–167. - PubMed
-
- Chan MK. Recent advances in heme-protein sensors. Curr Opin Chem Biol. 2001;5(2):216–222. - PubMed
-
- Olson JS, Phillips GN., Jr Kinetic pathways and barriers for ligand binding to myoglobin. J Biol Chem. 1996;271(30):17593–17596. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

