Mutations in Sensor 1 and Walker B in the bovine papillomavirus E1 initiator protein mimic the nucleotide-bound state
- PMID: 19939914
- PMCID: PMC2812389
- DOI: 10.1128/JVI.01756-09
Mutations in Sensor 1 and Walker B in the bovine papillomavirus E1 initiator protein mimic the nucleotide-bound state
Abstract
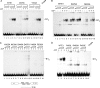
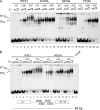
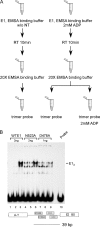
Viral replication initiator proteins are multifunctional proteins that utilize ATP binding and hydrolysis by their AAA+ modules for multiple functions in the replication of their viral genomes. These proteins are therefore of particular interest for understanding how AAA+ proteins carry out multiple ATP driven functions. We have performed a comprehensive mutational analysis of the residues involved in ATP binding and hydrolysis in the papillomavirus E1 initiator protein based on the recent structural data. Ten of the eleven residues that were targeted were defective for ATP hydrolysis, and seven of these were also defective for ATP binding. The three mutants that could still bind nucleotide represent the Walker B motif (D478 and D479) and Sensor 1 (N523), three residues that are in close proximity to each other and generally are considered to be involved in ATP hydrolysis. Surprisingly, however, two of these mutants, D478A and N523A, mimicked the nucleotide bound state and were capable of binding DNA in the absence of nucleotide. However, these mutants could not form the E1 double trimer in the absence of nucleotide, demonstrating that there are two qualitatively different consequences of ATP binding by E1, one that can be mimicked by D478A and N523A and one which cannot.
Figures


Similar articles
-
Transient viral DNA replication and repression of viral transcription are supported by the C-terminal domain of the bovine papillomavirus type 1 E1 protein.J Virol. 1998 Jan;72(1):796-801. doi: 10.1128/JVI.72.1.796-801.1998. J Virol. 1998. PMID: 9420289 Free PMC article.
-
Surface mutagenesis of the bovine papillomavirus E1 DNA binding domain reveals residues required for multiple functions related to DNA replication.J Virol. 2006 Aug;80(15):7491-9. doi: 10.1128/JVI.00435-06. J Virol. 2006. PMID: 16840329 Free PMC article.
-
Sequential and ordered assembly of E1 initiator complexes on the papillomavirus origin of DNA replication generates progressive structural changes related to melting.Mol Cell Biol. 2002 Nov;22(21):7712-20. doi: 10.1128/MCB.22.21.7712-7720.2002. Mol Cell Biol. 2002. PMID: 12370317 Free PMC article.
-
DNA-binding domain of bovine papillomavirus type 1 E1 helicase: structural and functional aspects.J Virol. 1993 Oct;67(10):6000-14. doi: 10.1128/JVI.67.10.6000-6014.1993. J Virol. 1993. PMID: 8396665 Free PMC article.
-
Bovine papillomavirus type 1 E1 and simian virus 40 large T antigen share regions of sequence similarity required for multiple functions.J Virol. 1997 Oct;71(10):7600-8. doi: 10.1128/JVI.71.10.7600-7608.1997. J Virol. 1997. PMID: 9311841 Free PMC article.
Cited by
-
A conserved regulatory module at the C terminus of the papillomavirus E1 helicase domain controls E1 helicase assembly.J Virol. 2015 Jan 15;89(2):1129-42. doi: 10.1128/JVI.01903-14. Epub 2014 Nov 5. J Virol. 2015. PMID: 25378487 Free PMC article.
-
The E1 proteins.Virology. 2013 Oct;445(1-2):35-56. doi: 10.1016/j.virol.2013.07.020. Epub 2013 Sep 10. Virology. 2013. PMID: 24029589 Free PMC article. Review.
-
Engagement of the ATR-dependent DNA damage response at the human papillomavirus 18 replication centers during the initial amplification.J Virol. 2013 Jan;87(2):951-64. doi: 10.1128/JVI.01943-12. Epub 2012 Nov 7. J Virol. 2013. PMID: 23135710 Free PMC article.
References
-
- Enemark, E. J., and L. Joshua-Tor. 2006. Mechanism of DNA translocation in a replicative hexameric helicase. Nature 442:270-275. - PubMed
-
- Gai, D., R. Zhao, D. Li, C. V. Finkielstein, and X. S. Chen. 2004. Mechanisms of conformational change for a replicative hexameric helicase of SV40 large tumor antigen. Cell 119:47-60. - PubMed
-
- Gorbalenya, A. E., E. V. Koonin, and Y. I. Wolf. 1990. A new superfamily of putative NTP-binding domains encoded by genomes of small DNA and RNA viruses. FEBS Lett. 262:145-148. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

