The process-inducing activity of transmembrane agrin requires follistatin-like domains
- PMID: 19940118
- PMCID: PMC2823450
- DOI: 10.1074/jbc.M109.039420
The process-inducing activity of transmembrane agrin requires follistatin-like domains
Abstract
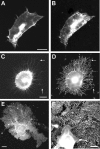
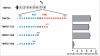
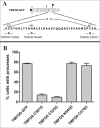
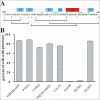
Clustering or overexpression of the transmembrane form of the extracellular matrix proteoglycan agrin in neurons results in the formation of numerous highly motile filopodia-like processes extending from axons and dendrites. Here we show that similar processes can be induced by overexpression of transmembrane-agrin in several non-neuronal cell lines. Mapping of the process-inducing activity in neurons and non-neuronal cells demonstrates that the cytoplasmic part of transmembrane agrin is dispensable and that the extracellular region is necessary for process formation. Site-directed mutagenesis reveals an essential role for the loop between beta-sheets 3 and 4 within the Kazal subdomain of the seventh follistatin-like domain of TM-agrin. An aspartic acid residue within this loop is critical for process formation. The seventh follistatin-like domain could be functionally replaced by the first and sixth but not by the eighth follistatin-like domain, demonstrating a functional redundancy among some follistatin-like domains of agrin. Moreover, a critical distance of the seventh follistatin-like domain to the plasma membrane appears to be required for process formation. These results demonstrate that different regions within the agrin protein are responsible for synapse formation at the neuromuscular junction and for process formation in central nervous system neurons and suggest a role for agrin's follistatin-like domains in the developing central nervous system.
Figures


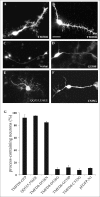
References
-
- Bezakova G., Ruegg M. A. (2003) Nat. Rev. Mol. Cell Biol. 4, 295–308 - PubMed
-
- Kröger S., Pfister H. (2009) Fut. Neurol. 4, 67–86
-
- Sanes J. R., Lichtman J. W. (2001) Nat. Rev. Neurosci. 2, 791–805 - PubMed
-
- Gautam M., Noakes P. G., Moscoso L., Rupp F., Scheller R. H., Merlie J. P., Sanes J. R. (1996) Cell 85, 525–535 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

