Cordycepin inhibits protein synthesis and cell adhesion through effects on signal transduction
- PMID: 19940154
- PMCID: PMC2807318
- DOI: 10.1074/jbc.M109.071159
Cordycepin inhibits protein synthesis and cell adhesion through effects on signal transduction
Abstract
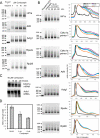
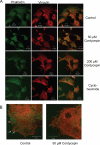
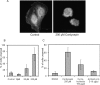
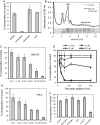
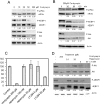
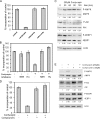
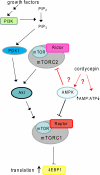
3'-Deoxyadenosine, also known as cordycepin, is a known polyadenylation inhibitor with a large spectrum of biological activities, including anti-proliferative, pro-apoptotic and anti-inflammatory effects. In this study we confirm that cordycepin reduces the length of poly(A) tails, with some mRNAs being much more sensitive than others. The low doses of cordycepin that cause poly(A) changes also reduce the proliferation of NIH3T3 fibroblasts. At higher doses of the drug we observed inhibition of cell attachment and a reduction of focal adhesions. Furthermore, we observed a strong inhibition of total protein synthesis that correlates with an inhibition of mammalian target of rapamycin (mTOR) signaling, as observed by reductions in Akt kinase and 4E-binding protein (4EBP) phosphorylation. In 4EBP knock-out cells, the effect of cordycepin on translation is strongly reduced, confirming the role of this modification. In addition, the AMP-activated kinase (AMPK) was shown to be activated. Inhibition of AMPK prevented translation repression by cordycepin and abolished 4EBP1 dephosphorylation, indicating that the effect of cordycepin on mTOR signaling and protein synthesis is mediated by AMPK activation. We conclude that many of the reported biological effects of cordycepin are likely to be due to its effects on mTOR and AMPK signaling.
Figures
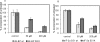
Similar articles
-
Cordycepin activates AMP-activated protein kinase (AMPK) via interaction with the γ1 subunit.J Cell Mol Med. 2014 Feb;18(2):293-304. doi: 10.1111/jcmm.12187. Epub 2013 Nov 28. J Cell Mol Med. 2014. PMID: 24286368 Free PMC article.
-
Cordycepin generally inhibits growth factor signal transduction in a systems pharmacology study.FEBS Lett. 2025 Feb;599(3):415-435. doi: 10.1002/1873-3468.15046. Epub 2024 Nov 7. FEBS Lett. 2025. PMID: 39508147 Free PMC article.
-
Inhibition of polyadenylation reduces inflammatory gene induction.RNA. 2012 Dec;18(12):2236-50. doi: 10.1261/rna.032391.112. Epub 2012 Nov 1. RNA. 2012. PMID: 23118416 Free PMC article.
-
Cordycepin and kinase inhibition in cancer.Drug Discov Today. 2023 Mar;28(3):103481. doi: 10.1016/j.drudis.2022.103481. Epub 2022 Dec 27. Drug Discov Today. 2023. PMID: 36584876 Review.
-
Application and research progress of cordycepin in the treatment of tumours (Review).Mol Med Rep. 2024 Sep;30(3):161. doi: 10.3892/mmr.2024.13285. Epub 2024 Jul 12. Mol Med Rep. 2024. PMID: 38994776 Free PMC article. Review.
Cited by
-
Traditional Chinese Medicine, Cordyceps, Related to Hepatoportal Sclerosis.ACG Case Rep J. 2023 Nov 29;10(12):e01206. doi: 10.14309/crj.0000000000001206. eCollection 2023 Dec. ACG Case Rep J. 2023. PMID: 38033616 Free PMC article.
-
Fluorescence Detection of Deoxyadenosine in Cordyceps spp. by Indicator Displacement Assay.Molecules. 2020 Apr 28;25(9):2045. doi: 10.3390/molecules25092045. Molecules. 2020. PMID: 32353945 Free PMC article.
-
Protective effects of cordycepin pretreatment against liver ischemia/reperfusion injury in mice.Immun Inflamm Dis. 2023 Mar;11(3):e792. doi: 10.1002/iid3.792. Immun Inflamm Dis. 2023. PMID: 36988254 Free PMC article.
-
The polyadenylation inhibitor cordycepin reduces pain, inflammation and joint pathology in rodent models of osteoarthritis.Sci Rep. 2019 Mar 18;9(1):4696. doi: 10.1038/s41598-019-41140-1. Sci Rep. 2019. PMID: 30886197 Free PMC article.
-
Pharmacological and therapeutic potential of Cordyceps with special reference to Cordycepin.3 Biotech. 2014 Feb;4(1):1-12. doi: 10.1007/s13205-013-0121-9. Epub 2013 Feb 19. 3 Biotech. 2014. PMID: 28324458 Free PMC article.
References
-
- Yang F. Q., Li S. P. (2008) J. Pharm. Biomed. Anal. 48, 231–235 - PubMed
-
- Ng T. B., Wang H. X. (2005) J. Pharm. Pharmacol. 57, 1509–1519 - PubMed
-
- Chang W., Lim S., Song H., Song B. W., Kim H. J., Cha M. J., Sung J. M., Kim T. W., Hwang K. C. (2008) Eur. J. Pharmacol. 597, 64–69 - PubMed
-
- Shi P., Huang Z., Tan X., Chen G. (2008) Methods Find. Exp. Clin. Pharmacol. 30, 347–353 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

