Myristylation and palmitylation of HSV-1 UL11 are not essential for its function
- PMID: 19944438
- PMCID: PMC2813933
- DOI: 10.1016/j.virol.2009.10.046
Myristylation and palmitylation of HSV-1 UL11 are not essential for its function
Abstract
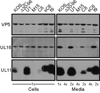
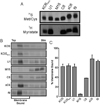
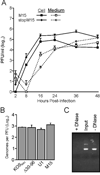
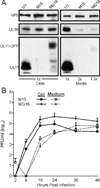
All herpesviruses encode a homolog of the herpes simplex virus type-1 UL11 tegument protein. Deletion of UL11 disrupts virus envelopment, causes capsid accumulation within the cytoplasm, and reduces virus release. UL11 requires acylation with myristate and palmitate for membrane binding, lipid raft trafficking, and accumulation at the site of virus envelopment. Thus, it was predicted that acylation of UL11 would be necessary for efficient virion production, similar to HIV-1 Gag which requires myristylation for virus production. Accordingly, recombinant viruses were created to express UL11 derivatives that are not acylated, are partially acylated, or contain foreign acylation signals. Unexpectedly, the non-acylated UL11 rescued some growth defects of a UL11-null mutant, even though the unmodified protein was unstable. Furthermore, a myristylated and palmitylated chimera did not fully rescue the null virus. These results suggest that UL11 maintains some function(s) when not membrane-bound, and the sequence context of the acylations is important for UL11 function.
Copyright 2009 Elsevier Inc. All rights reserved.
Figures



References
-
- Desai P, DeLuca NA, Person S. Herpes simplex virus type 1 VP26 is not essential for replication in cell culture but influences production of infectious virus in the nervous system of infected mice. Virology. 1998;247:115–124. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

