The microwell control of embryoid body size in order to regulate cardiac differentiation of human embryonic stem cells
- PMID: 19945747
- PMCID: PMC2813988
- DOI: 10.1016/j.biomaterials.2009.11.033
The microwell control of embryoid body size in order to regulate cardiac differentiation of human embryonic stem cells
Abstract
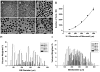
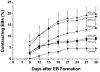
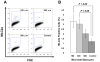
The differentiation of human embryonic stem cells (hESCs) into cardiomyocytes (CMs) using embryoid bodies (EBs) is relatively inefficient and highly variable. Formation of EBs using standard enzymatic disaggregation techniques results in a wide range of sizes and geometries of EBs. Use of a 3-D cuboidal microwell system to culture hESCs in colonies of defined dimensions, 100-500 microm in lateral dimensions and 120 microm in depth, enabled formation of more uniform-sized EBs. The 300 microm microwells produced highest percentage of contracting EBs, but flow cytometry for myosin light chain 2A (MLC2a) expressing cells revealed a similar percentage (approximately 3%) of cardiomyocytes formed in EBs from 100 microm to 300 microm microwells. These data, and immunolabeling with anti-MF20 and MLC2a, suggest that the smaller EBs are less likely to form contracting EBs, but those contracting EBs are relatively enriched in cardiomyocytes compared to larger EB sizes where CMs make up a proportionately smaller fraction of the total cells. We conclude that microwell-engineered EB size regulates cardiogenesis and can be used for more efficient and reproducible formation of hESC-CMs needed for research and therapeutic applications.
(c) 2009 Elsevier Ltd. All rights reserved.
Figures

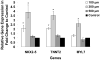
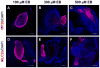
References
-
- Odorico JS, Kaufman DS, Thomson JA. Multilineage differentiation from human embryonic stem cell lines. Stem Cells. 2001;19:193–204. - PubMed
-
- Watt FM, Hogan BL. Out of Eden: stem cells and their niches. Science. 2000;287:1427–30. - PubMed
-
- Spradling A, Drummond-Barbosa D, Kai T. Stem cells find their niche. Nature. 2001;414:98–104. - PubMed
-
- Streuli C. Extracellular matrix remodelling and cellular differentiation. Curr Opin Cell Biol. 1999;11:634–40. - PubMed
-
- Mummery C, Ward-van Oostwaard D, Doevendans P, Spijker R, van den BS, Hassink R, et al. Differentiation of human embryonic stem cells to cardiomyocytes: role of coculture with visceral endoderm-like cells. Circulation. 2003;107:2733–40. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

