Arabidopsis N-MYC DOWNREGULATED-LIKE1, a positive regulator of auxin transport in a G protein-mediated pathway
- PMID: 19948787
- PMCID: PMC2798320
- DOI: 10.1105/tpc.109.065557
Arabidopsis N-MYC DOWNREGULATED-LIKE1, a positive regulator of auxin transport in a G protein-mediated pathway
Abstract
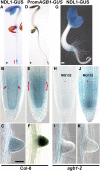
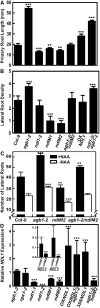
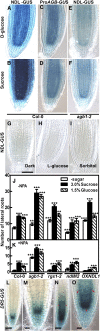
Root architecture results from coordinated cell division and expansion in spatially distinct cells of the root and is established and maintained by gradients of auxin and nutrients such as sugars. Auxin is transported acropetally through the root within the central stele and then, upon reaching the root apex, auxin is transported basipetally through the outer cortical and epidermal cells. The two Gbetagamma dimers of the Arabidopsis thaliana heterotrimeric G protein complex are differentially localized to the central and cortical tissues of the Arabidopsis roots. A null mutation in either the single beta (AGB1) or the two gamma (AGG1 and AGG2) subunits confers phenotypes that disrupt the proper architecture of Arabidopsis roots and are consistent with altered auxin transport. Here, we describe an evolutionarily conserved interaction between AGB1/AGG dimers and a protein designated N-MYC DOWNREGULATED-LIKE1 (NDL1). The Arabidopsis genome encodes two homologs of NDL1 (NDL2 and NDL3), which also interact with AGB1/AGG1 and AGB1/AGG2 dimers. We show that NDL proteins act in a signaling pathway that modulates root auxin transport and auxin gradients in part by affecting the levels of at least two auxin transport facilitators. Reduction of NDL family gene expression and overexpression of NDL1 alter root architecture, auxin transport, and auxin maxima. AGB1, auxin, and sugars are required for NDL1 protein stability in regions of the root where auxin gradients are established; thus, the signaling mechanism contains feedback loops.
Figures





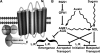
References
-
- Akgoz, M., Azpiazu, I., Kalyanaraman, V., and Gautam, N. (2002). Role of the G protein gamma subunit in beta gamma complex modulation of phospholipase Cβ function. J. Biol. Chem. 277 19573–19578. - PubMed
-
- Benjamins, R., Malenica, N., and Luschnig, C. (2005). Regulating the regulator: the control of auxin transport. Bioessays 27 1246–1255. - PubMed
-
- Benjamins, R., and Scheres, B. (2008). Auxin: The looping star in plant development. Annu. Rev. Plant Biol. 59 443–465. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

