Genome sequence of the Fleming strain of Micrococcus luteus, a simple free-living actinobacterium
- PMID: 19948807
- PMCID: PMC2812450
- DOI: 10.1128/JB.01254-09
Genome sequence of the Fleming strain of Micrococcus luteus, a simple free-living actinobacterium
Abstract
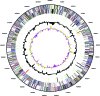
Micrococcus luteus (NCTC2665, "Fleming strain") has one of the smallest genomes of free-living actinobacteria sequenced to date, comprising a single circular chromosome of 2,501,097 bp (G+C content, 73%) predicted to encode 2,403 proteins. The genome shows extensive synteny with that of the closely related organism, Kocuria rhizophila, from which it was taxonomically separated relatively recently. Despite its small size, the genome harbors 73 insertion sequence (IS) elements, almost all of which are closely related to elements found in other actinobacteria. An IS element is inserted into the rrs gene of one of only two rrn operons found in M. luteus. The genome encodes only four sigma factors and 14 response regulators, a finding indicative of adaptation to a rather strict ecological niche (mammalian skin). The high sensitivity of M. luteus to beta-lactam antibiotics may result from the presence of a reduced set of penicillin-binding proteins and the absence of a wblC gene, which plays an important role in the antibiotic resistance in other actinobacteria. Consistent with the restricted range of compounds it can use as a sole source of carbon for energy and growth, M. luteus has a minimal complement of genes concerned with carbohydrate transport and metabolism and its inability to utilize glucose as a sole carbon source may be due to the apparent absence of a gene encoding glucokinase. Uniquely among characterized bacteria, M. luteus appears to be able to metabolize glycogen only via trehalose and to make trehalose only via glycogen. It has very few genes associated with secondary metabolism. In contrast to most other actinobacteria, M. luteus encodes only one resuscitation-promoting factor (Rpf) required for emergence from dormancy, and its complement of other dormancy-related proteins is also much reduced. M. luteus is capable of long-chain alkene biosynthesis, which is of interest for advanced biofuel production; a three-gene cluster essential for this metabolism has been identified in the genome.
Figures


Similar articles
-
Comparative genomics reveals broad genetic diversity, extensive recombination and nascent ecological adaptation in Micrococcus luteus.BMC Genomics. 2021 Feb 18;22(1):124. doi: 10.1186/s12864-021-07432-5. BMC Genomics. 2021. PMID: 33602135 Free PMC article.
-
Structural changes and cellular localization of resuscitation-promoting factor in environmental isolates of Micrococcus luteus.Microb Ecol. 2010 Feb;59(2):296-310. doi: 10.1007/s00248-009-9573-1. Microb Ecol. 2010. PMID: 19730766
-
Comparative metabolic capabilities for Micrococcus luteus NCTC 2665, the "Fleming" strain, and actinobacteria.Biotechnol Bioeng. 2011 Nov;108(11):2770-5. doi: 10.1002/bit.23212. Epub 2011 Jun 9. Biotechnol Bioeng. 2011. PMID: 21618466
-
A Proteomic Signature of Dormancy in the Actinobacterium Micrococcus luteus.J Bacteriol. 2017 Jun 27;199(14):e00206-17. doi: 10.1128/JB.00206-17. Print 2017 Jul 15. J Bacteriol. 2017. PMID: 28484042 Free PMC article.
-
Penicillin-Binding Proteins, β-Lactamases, and β-Lactamase Inhibitors in β-Lactam-Producing Actinobacteria: Self-Resistance Mechanisms.Int J Mol Sci. 2022 May 18;23(10):5662. doi: 10.3390/ijms23105662. Int J Mol Sci. 2022. PMID: 35628478 Free PMC article. Review.
Cited by
-
Beyond asexual development: modifications in the gene expression profile caused by the absence of the Aspergillus nidulans transcription factor FlbB.Genetics. 2015 Apr;199(4):1127-42. doi: 10.1534/genetics.115.174342. Epub 2015 Feb 20. Genetics. 2015. PMID: 25701285 Free PMC article.
-
Negative Correlation between Lipid Content and Antibiotic Activity in Streptomyces: General Rule and Exceptions.Antibiotics (Basel). 2020 May 26;9(6):280. doi: 10.3390/antibiotics9060280. Antibiotics (Basel). 2020. PMID: 32466356 Free PMC article.
-
Draft Genome Sequence of Micrococcus sp. Strain MS-AsIII-49, an Arsenate-Reducing Isolate from Tropical Metal-Rich Sediment.Genome Announc. 2015 Apr 16;3(2):e00122-15. doi: 10.1128/genomeA.00122-15. Genome Announc. 2015. PMID: 25883272 Free PMC article.
-
Heterotrophic microflora of highly alkaline (pH > 13) brown mud disposal site drainage water near Ziar nad Hronom (Banska Bystrica region, Slovakia).Environ Sci Pollut Res Int. 2016 Mar;23(5):4199-206. doi: 10.1007/s11356-015-4842-7. Epub 2015 Jun 17. Environ Sci Pollut Res Int. 2016. PMID: 26077319
-
Comparative genomics reveals broad genetic diversity, extensive recombination and nascent ecological adaptation in Micrococcus luteus.BMC Genomics. 2021 Feb 18;22(1):124. doi: 10.1186/s12864-021-07432-5. BMC Genomics. 2021. PMID: 33602135 Free PMC article.
References
-
- Albro, P. W., and J. C. Dittmer. 1969. The biochemistry of long-chain, nonisoprenoid hydrocarbons. I. Characterization of the hydrocarbons of Sarcina lutea and the isolation of possible intermediates of biosynthesis. Biochemistry 8:394-404. - PubMed
-
- Anwar, M., T. H. Khan, J. Prebble, and P. F. Zagalsky. 1977. Membrane-bound carotenoid in Micrococcus luteus protects naphthoquinone from photodynamic action. Nature 270:538-540. - PubMed
-
- Artsatbanov, V., G. V. Tikhonova, and D. N. Ostrovskii. 1983. Generation of membrane potential by aerobic bacteria Micrococcus lysodeikticus. Correlation between coupled and uncoupled respiration. Biokhimiia 48:1568-1579. (In Russian.) - PubMed
Publication types
MeSH terms
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

