Ube2j2 ubiquitinates hydroxylated amino acids on ER-associated degradation substrates
- PMID: 19951915
- PMCID: PMC2806592
- DOI: 10.1083/jcb.200908036
Ube2j2 ubiquitinates hydroxylated amino acids on ER-associated degradation substrates
Abstract
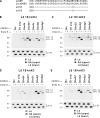
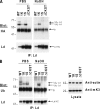
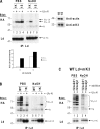
Ubiquitin (Ub) modification of proteins plays a prominent role in the regulation of multiple cell processes, including endoplasmic reticulum-associated degradation (ERAD). Until recently, ubiquitination of substrates was thought to occur only via isopeptide bonds, typically to lysine residues. Several recent studies suggest that Ub can also be coupled to nonlysine residues by ester/thiolester bonds; however, the molecular basis for these novel modifications remains elusive. To probe the mechanism and importance of nonlysine ubiquitination, we have studied the viral ligase murine K3 (mK3), which facilitates the polyubiquitination of hydroxylated amino acids serine/threonine on its ERAD substrate. In this paper, we identify Ube2j2 as the primary cellular E2 recruited by the mK3 ligase, and this E2-E3 pair is capable of conjugating Ub on lysine or serine residues of substrates. However, surprisingly, Ube2j2-mK3 preferentially promotes ubiquitination of hydroxylated amino acids via ester bonds even when lysine residues are present on wild-type substrates, thus establishing physiological relevance of this novel ubiquitination strategy.
Figures
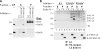


Similar articles
-
Role of the RING-CH domain of viral ligase mK3 in ubiquitination of non-lysine and lysine MHC I residues.Traffic. 2009 Sep;10(9):1301-17. doi: 10.1111/j.1600-0854.2009.00946.x. Epub 2009 May 27. Traffic. 2009. PMID: 19531064
-
Ubiquitination of serine, threonine, or lysine residues on the cytoplasmic tail can induce ERAD of MHC-I by viral E3 ligase mK3.J Cell Biol. 2007 May 21;177(4):613-24. doi: 10.1083/jcb.200611063. Epub 2007 May 14. J Cell Biol. 2007. PMID: 17502423 Free PMC article.
-
Molecular basis for lysine specificity in the yeast ubiquitin-conjugating enzyme Cdc34.Mol Cell Biol. 2010 May;30(10):2316-29. doi: 10.1128/MCB.01094-09. Epub 2010 Mar 1. Mol Cell Biol. 2010. PMID: 20194622 Free PMC article.
-
Ubiquitination of substrates by esterification.Traffic. 2012 Jan;13(1):19-24. doi: 10.1111/j.1600-0854.2011.01269.x. Epub 2011 Sep 13. Traffic. 2012. PMID: 21883762 Free PMC article. Review.
-
Diverse roles of the E2/E3 hybrid enzyme UBE2O in the regulation of protein ubiquitination, cellular functions, and disease onset.FEBS J. 2019 Jun;286(11):2018-2034. doi: 10.1111/febs.14708. Epub 2018 Dec 4. FEBS J. 2019. PMID: 30468556 Review.
Cited by
-
Biomarker discovery in attention deficit hyperactivity disorder: RNA sequencing of whole blood in discordant twin and case-controlled cohorts.BMC Med Genomics. 2020 Oct 28;13(1):160. doi: 10.1186/s12920-020-00808-8. BMC Med Genomics. 2020. PMID: 33115496 Free PMC article.
-
E3 ubiquitin ligases in ErbB receptor quantity control.Semin Cell Dev Biol. 2010 Dec;21(9):936-43. doi: 10.1016/j.semcdb.2010.09.006. Epub 2010 Sep 22. Semin Cell Dev Biol. 2010. PMID: 20868762 Free PMC article. Review.
-
How ubiquitin functions with ESCRTs.Traffic. 2011 Oct;12(10):1306-17. doi: 10.1111/j.1600-0854.2011.01242.x. Epub 2011 Jul 27. Traffic. 2011. PMID: 21722280 Free PMC article. Review.
-
Disposing of misfolded ER proteins: A troubled substrate's way out of the ER.Mol Cell Endocrinol. 2020 Jan 15;500:110630. doi: 10.1016/j.mce.2019.110630. Epub 2019 Oct 24. Mol Cell Endocrinol. 2020. PMID: 31669350 Free PMC article. Review.
-
Ubiquitylation of an ERAD substrate occurs on multiple types of amino acids.Mol Cell. 2010 Dec 22;40(6):917-26. doi: 10.1016/j.molcel.2010.11.033. Mol Cell. 2010. PMID: 21172657 Free PMC article.
References
-
- Arteaga M.F., Wang L., Ravid T., Hochstrasser M., Canessa C.M. 2006. An amphipathic helix targets serum and glucocorticoid-induced kinase 1 to the endoplasmic reticulum-associated ubiquitin-conjugation machinery. Proc. Natl. Acad. Sci. USA. 103:11178–11183 10.1073/pnas.0604816103 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

