Homology-dependent interactions determine the order of strand exchange by IntDOT recombinase
- PMID: 19952068
- PMCID: PMC2817482
- DOI: 10.1093/nar/gkp927
Homology-dependent interactions determine the order of strand exchange by IntDOT recombinase
Abstract
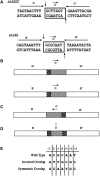
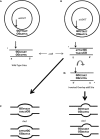
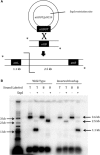
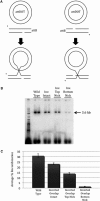
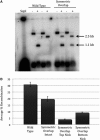
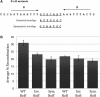
The Bacteroides conjugative transposon CTnDOT encodes an integrase, IntDOT, which is a member of the tyrosine recombinase family. Other members of this group share a strict requirement for sequence identity within the region of strand exchange, called the overlap region. Tyrosine recombinases catalyze recombination by making an initial cleavage, strand exchange and ligation, followed by strand swapping isomerization requiring sequence identity in the overlap region, followed by the second cleavage, strand exchange and ligation. IntDOT is of particular interest because it has been shown to utilize a three-step mechanism: a sequence identity-dependent initial strand exchange that requires two base pairs of complementary DNA at the site of cleavage; a sequence identity-independent strand swapping isomerization, followed by a sequence identity-independent cleavage, strand exchange and ligation. In addition to the sequence identity requirement in the overlap region, Lambda Int interactions with arm-type sites dictate the order of strand exchange regardless of the orientation of the overlap region. Although IntDOT has an arm-binding domain, we show here that the location of sequence identity within the overlap region dictates where the initial cleavage takes place and that IntDOT can recombine substrates containing mismatches in the overlap region so long as a single base of sequence identity exists at the site of initial cleavage.
Figures


Similar articles
-
Resolution of Mismatched Overlap Holliday Junction Intermediates by the Tyrosine Recombinase IntDOT.J Bacteriol. 2017 Apr 25;199(10):e00873-16. doi: 10.1128/JB.00873-16. Print 2017 May 15. J Bacteriol. 2017. PMID: 28242723 Free PMC article.
-
The Integration and Excision of CTnDOT.Microbiol Spectr. 2015 Apr;3(2):MDNA3-0020-2014. doi: 10.1128/microbiolspec.MDNA3-0020-2014. Microbiol Spectr. 2015. PMID: 26104696 Free PMC article. Review.
-
CTnDOT integrase performs ordered homology-dependent and homology-independent strand exchanges.Nucleic Acids Res. 2007;35(17):5861-73. doi: 10.1093/nar/gkm637. Epub 2007 Aug 24. Nucleic Acids Res. 2007. PMID: 17720706 Free PMC article.
-
Characterization of a conjugative transposon integrase, IntDOT.Mol Microbiol. 2006 Jun;60(5):1228-40. doi: 10.1111/j.1365-2958.2006.05164.x. Mol Microbiol. 2006. PMID: 16689798
-
Challenging a paradigm: the role of DNA homology in tyrosine recombinase reactions.Microbiol Mol Biol Rev. 2009 Jun;73(2):300-9. doi: 10.1128/MMBR.00038-08. Microbiol Mol Biol Rev. 2009. PMID: 19487729 Free PMC article. Review.
Cited by
-
Evolutionary dynamics and genomic features of the Elizabethkingia anophelis 2015 to 2016 Wisconsin outbreak strain.Nat Commun. 2017 May 24;8:15483. doi: 10.1038/ncomms15483. Nat Commun. 2017. PMID: 28537263 Free PMC article.
-
Resolution of Mismatched Overlap Holliday Junction Intermediates by the Tyrosine Recombinase IntDOT.J Bacteriol. 2017 Apr 25;199(10):e00873-16. doi: 10.1128/JB.00873-16. Print 2017 May 15. J Bacteriol. 2017. PMID: 28242723 Free PMC article.
-
The Integration and Excision of CTnDOT.Microbiol Spectr. 2015 Apr;3(2):MDNA3-0020-2014. doi: 10.1128/microbiolspec.MDNA3-0020-2014. Microbiol Spectr. 2015. PMID: 26104696 Free PMC article. Review.
-
Horizontal gene transfers with or without cell fusions in all categories of the living matter.Adv Exp Med Biol. 2011;714:5-89. doi: 10.1007/978-94-007-0782-5_2. Adv Exp Med Biol. 2011. PMID: 21506007 Free PMC article. Review.
-
Interactions of the excision proteins of CTnDOT in the attR intasome.Plasmid. 2013 Sep;70(2):190-200. doi: 10.1016/j.plasmid.2013.03.009. Epub 2013 Apr 17. Plasmid. 2013. PMID: 23603449 Free PMC article.

