Stu1 inversely regulates kinetochore capture and spindle stability
- PMID: 19952112
- PMCID: PMC2788326
- DOI: 10.1101/gad.541309
Stu1 inversely regulates kinetochore capture and spindle stability
Abstract
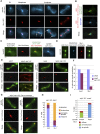
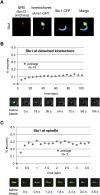
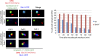
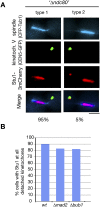
The Saccharomyces cerevisiae CLASP (CLIP-associated protein) Stu1 is essential for the establishment and maintenance of the mitotic spindle. Furthermore, Stu1 localizes to kinetochores. Here we show that, in prometaphase, Stu1 assembles in an Ndc80-dependent manner exclusively at kinetochores that are not attached to microtubules. Stu1 relocates to microtubules when a captured kinetochore reaches a spindle pole. This relocation does not depend on kinetochore biorientation, but requires a functional DASH complex. Stu1 at detached kinetochores facilitates kinetochore capturing. Furthermore, since most of the nuclear Stu1 is sequestered by one or a few detached kinetochores, the presence of detached kinetochores prevents Stu1 from localizing the spindle, and therefore from stabilizing the spindle. Thus, the sequestering of Stu1 by detached kinetochores serves as a checkpoint that keeps spindle poles in close proximity until all kinetochores are captured. This is likely to facilitate kinetochore biorientation. In agreement with the findings described above, a kinetochore mutant (okp1-52) that fails to release Stu1 from the kinetochore displays a severe spindle defect upon spindle pole body separation, and this defect can be rescued by destroying the okp1-52 kinetochore.
Figures



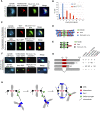
Similar articles
-
Unattached kinetochores drive their own capturing by sequestering a CLASP.Nat Commun. 2018 Feb 28;9(1):886. doi: 10.1038/s41467-018-03108-z. Nat Commun. 2018. PMID: 29491436 Free PMC article.
-
The composition, functions, and regulation of the budding yeast kinetochore.Genetics. 2013 Aug;194(4):817-46. doi: 10.1534/genetics.112.145276. Genetics. 2013. PMID: 23908374 Free PMC article. Review.
-
A TOGL domain specifically targets yeast CLASP to kinetochores to stabilize kinetochore microtubules.J Cell Biol. 2014 May 26;205(4):555-71. doi: 10.1083/jcb.201310018. J Cell Biol. 2014. PMID: 24862575 Free PMC article.
-
Molecular mechanisms of microtubule-dependent kinetochore transport toward spindle poles.J Cell Biol. 2007 Jul 16;178(2):269-81. doi: 10.1083/jcb.200702141. Epub 2007 Jul 9. J Cell Biol. 2007. PMID: 17620411 Free PMC article.
-
The Ndc80 complex: hub of kinetochore activity.FEBS Lett. 2007 Jun 19;581(15):2862-9. doi: 10.1016/j.febslet.2007.05.012. Epub 2007 May 11. FEBS Lett. 2007. PMID: 17521635 Review.
Cited by
-
GTP regulates the microtubule nucleation activity of γ-tubulin.Nat Cell Biol. 2013 Nov;15(11):1317-27. doi: 10.1038/ncb2863. Epub 2013 Oct 27. Nat Cell Biol. 2013. PMID: 24161932
-
Unattached kinetochores drive their own capturing by sequestering a CLASP.Nat Commun. 2018 Feb 28;9(1):886. doi: 10.1038/s41467-018-03108-z. Nat Commun. 2018. PMID: 29491436 Free PMC article.
-
The composition, functions, and regulation of the budding yeast kinetochore.Genetics. 2013 Aug;194(4):817-46. doi: 10.1534/genetics.112.145276. Genetics. 2013. PMID: 23908374 Free PMC article. Review.
-
chTOG is a conserved mitotic error correction factor.Elife. 2020 Dec 30;9:e61773. doi: 10.7554/eLife.61773. Elife. 2020. PMID: 33377866 Free PMC article.
-
Slk19 clusters kinetochores and facilitates chromosome bipolar attachment.Mol Biol Cell. 2013 Mar;24(5):566-77. doi: 10.1091/mbc.E12-07-0552. Epub 2013 Jan 2. Mol Biol Cell. 2013. PMID: 23283988 Free PMC article.
References
-
- Akhmanova A, Hoogenraad CC. Microtubule plus-end-tracking proteins: Mechanisms and functions. Curr Opin Cell Biol. 2005;17:47–54. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
