Regulation of class A scavenger receptor-mediated cell adhesion and surface localization by PI3K: identification of a regulatory cytoplasmic motif
- PMID: 19952357
- PMCID: PMC2830124
- DOI: 10.1189/jlb.0509318
Regulation of class A scavenger receptor-mediated cell adhesion and surface localization by PI3K: identification of a regulatory cytoplasmic motif
Abstract
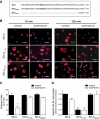
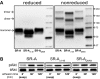
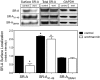
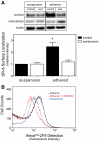
The importance of cytoplasmic motifs in differentially regulating SR-A function was demonstrated by deleting the first 49 cytoplasmic aa (SR-A(Delta1-49)), which abolished SR-A-mediated ligand internalization without reducing cell adhesion. To identify additional cytoplasmic motifs within the first 49 aa that regulate SR-A function, the acidic residues in a conserved motif (EDAD) were changed to their amide derivatives (SR-A(QNAN)). The function and regulation of SR-A(QNAN) were compared with that of SR-A(Delta1-49) and SR-A in transfected HEK-293 cells. Blocking PI3K activation inhibited SR-A, but not SR-A(Delta1-49)- or SR-A(QNAN)-mediated cell adhesion. Although deleting (SR-A(Delta1-49)) or mutating (SR-A(QNAN)) the EDAD motif abolished the PI3K sensitivity of SR-A-mediated cell adhesion, these mutations did not affect ligand internalization or PI3K activation during cell adhesion. To define the mechanism by which PI3K regulates SR-A-mediated cell adhesion, the cellular localization of wild-type and mutant SR-A was examined. PI3K inhibition reduced surface localization of SR-A but not of SR-A(Delta1-49) or SR-A(QNAN). The regulation of SR-A surface localization by PI3K was confirmed in peritoneal macrophages, which endogenously express SR-A. Together, these results suggest a pathway in which SR-A binding to an immobilized ligand activates PI3K to recruit more receptor to the plasma membrane and enhances cell adhesion.
Figures
Similar articles
-
Lipid rafts couple class A scavenger receptors to phospholipase A2 activation during macrophage adhesion.J Leukoc Biol. 2014 Nov;96(5):873-81. doi: 10.1189/jlb.2A0414-214R. Epub 2014 Jul 28. J Leukoc Biol. 2014. PMID: 25070949 Free PMC article.
-
Class A scavenger receptor-mediated adhesion and internalization require distinct cytoplasmic domains.J Biol Chem. 2003 Sep 5;278(36):34219-25. doi: 10.1074/jbc.M303465200. Epub 2003 Jun 20. J Biol Chem. 2003. PMID: 12819208
-
The di-leucine motif contributes to class a scavenger receptor-mediated internalization of acetylated lipoproteins.Arterioscler Thromb Vasc Biol. 2006 Jun;26(6):1317-22. doi: 10.1161/01.ATV.0000220171.50282.0c. Epub 2006 Mar 30. Arterioscler Thromb Vasc Biol. 2006. PMID: 16574888
-
[The role of the class A scavenger receptors, SR-A and MARCO, in the immune system. Part 1. The structure of receptors, their ligand binding repertoires and ability to initiate intracellular signaling].Postepy Hig Med Dosw (Online). 2012 Feb 29;66:104-19. doi: 10.5604/17322693.984079. Postepy Hig Med Dosw (Online). 2012. PMID: 22470185 Review. Polish.
-
Class A1 scavenger receptors in cardiovascular diseases.Br J Pharmacol. 2015 Dec;172(23):5523-30. doi: 10.1111/bph.13105. Epub 2015 Mar 27. Br J Pharmacol. 2015. PMID: 25651870 Free PMC article. Review.
Cited by
-
Cysteine-rich domain of scavenger receptor AI modulates the efficacy of surface targeting and mediates oligomeric Aβ internalization.J Biomed Sci. 2013 Aug 2;20(1):54. doi: 10.1186/1423-0127-20-54. J Biomed Sci. 2013. PMID: 23915271 Free PMC article.
-
Scavenger receptor-A (CD204): a two-edged sword in health and disease.Crit Rev Immunol. 2014;34(3):241-61. doi: 10.1615/critrevimmunol.2014010267. Crit Rev Immunol. 2014. PMID: 24941076 Free PMC article. Review.
-
Tumor-Associated CD204-Positive Macrophage Is a Prognostic Marker in Clinical Stage I Lung Adenocarcinoma.Biomed Res Int. 2018 Apr 16;2018:8459193. doi: 10.1155/2018/8459193. eCollection 2018. Biomed Res Int. 2018. PMID: 29850577 Free PMC article.
-
GRP78 inhibits macrophage adhesion via SR-A.J Biomed Res. 2014 Jul;28(4):269-74. doi: 10.7555/JBR.28.20130054. Epub 2014 Mar 20. J Biomed Res. 2014. PMID: 25050110 Free PMC article.
-
Scavenger receptor A1 participates in uptake of Leptospira interrogans serovar Autumnalis strain 56606v and inflammation in mouse macrophages.Emerg Microbes Infect. 2021 Dec;10(1):939-953. doi: 10.1080/22221751.2021.1925160. Emerg Microbes Infect. 2021. PMID: 33929941 Free PMC article.
References
-
- Pluddemann A, Neyen C, Gordon S. Macrophage scavenger receptors and host-derived ligands. Methods. 2007;43:207–217. - PubMed
-
- Platt N, Haworth R, Darley L, Gordon S. The many roles of the class A macrophage scavenger receptor. Int Rev Cytol. 2002;212:1–40. - PubMed
-
- Gowen B B, Borg T K, Ghaffar A, Mayer E P. Selective adhesion of macrophages to denatured forms of type I collagen is mediated by scavenger receptors. Matrix Biol. 2000;19:61–71. - PubMed
-
- Gowen B B, Borg T K, Ghaffar A, Mayer E P. The collagenous domain of class A scavenger receptors is involved in macrophage adhesion to collagens. J Leukoc Biol. 2001;69:575–582. - PubMed
-
- Santiago-Garcia J, Kodama T, Pitas R E. The class A scavenger receptor binds to proteoglycans and mediates adhesion of macrophages to the extracellular matrix. J Biol Chem. 2003;278:6942–6946. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

