Repressive and non-repressive chromatin at native telomeres in Saccharomyces cerevisiae
- PMID: 19954519
- PMCID: PMC3225887
- DOI: 10.1186/1756-8935-2-18
Repressive and non-repressive chromatin at native telomeres in Saccharomyces cerevisiae
Abstract
Background: In Saccharomyces cerevisiae genes that are located close to a telomere can become transcriptionally repressed by an epigenetic process known as telomere position effect. There is large variation in the level of the telomere position effect among telomeres, with many native ends exhibiting little repression.
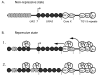
Results: Chromatin analysis, using microccocal nuclease and indirect end labelling, reveals distinct patterns for ends with different silencing states. Differences were observed in the promoter accessibility of a subtelomeric reporter gene and a characteristic array of phased nucleosomes was observed on the centromere proximal side of core X at a repressive end. The silent information regulator proteins 2 - 4, the yKu heterodimer and the subtelomeric core X element are all required for the maintenance of the chromatin structure of repressive ends. However, gene deletions of particular histone modification proteins can eliminate the silencing without the disruption of this chromatin structure.
Conclusion: Our data identifies chromatin features that correlate with the silencing state and indicate that an array of phased nucleosomes is not sufficient for full repression.
Figures





References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

