Tumor Classification Using High-Order Gene Expression Profiles Based on Multilinear ICA
- PMID: 19956422
- PMCID: PMC2778791
- DOI: 10.1155/2009/926450
Tumor Classification Using High-Order Gene Expression Profiles Based on Multilinear ICA
Abstract
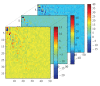
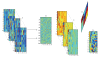
Motivation. Independent Components Analysis (ICA) maximizes the statistical independence of the representational components of a training gene expression profiles (GEP) ensemble, but it cannot distinguish relations between the different factors, or different modes, and it is not available to high-order GEP Data Mining. In order to generalize ICA, we introduce Multilinear-ICA and apply it to tumor classification using high order GEP. Firstly, we introduce the basis conceptions and operations of tensor and recommend Support Vector Machine (SVM) classifier and Multilinear-ICA. Secondly, the higher score genes of original high order GEP are selected by using t-statistics and tabulate tensors. Thirdly, the tensors are performed by Multilinear-ICA. Finally, the SVM is used to classify the tumor subtypes. Results. To show the validity of the proposed method, we apply it to tumor classification using high order GEP. Though we only use three datasets, the experimental results show that the method is effective and feasible. Through this survey, we hope to gain some insight into the problem of high order GEP tumor classification, in aid of further developing more effective tumor classification algorithms.
Figures



Similar articles
-
SGL-SVM: A novel method for tumor classification via support vector machine with sparse group Lasso.J Theor Biol. 2020 Feb 7;486:110098. doi: 10.1016/j.jtbi.2019.110098. Epub 2019 Nov 28. J Theor Biol. 2020. PMID: 31786183
-
Hybrid Feature Selection Algorithm mRMR-ICA for Cancer Classification from Microarray Gene Expression Data.Comb Chem High Throughput Screen. 2018;21(6):420-430. doi: 10.2174/1386207321666180601074349. Comb Chem High Throughput Screen. 2018. PMID: 29852866
-
Gene expression data classification using consensus independent component analysis.Genomics Proteomics Bioinformatics. 2008 Jun;6(2):74-82. doi: 10.1016/S1672-0229(08)60022-4. Genomics Proteomics Bioinformatics. 2008. PMID: 18973863 Free PMC article.
-
A fuzzy based feature selection from independent component subspace for machine learning classification of microarray data.Genom Data. 2016 Feb 23;8:4-15. doi: 10.1016/j.gdata.2016.02.012. eCollection 2016 Jun. Genom Data. 2016. PMID: 27081632 Free PMC article.
-
[Hyperspectral remote sensing image classification based on ICA and SVM algorithm].Guang Pu Xue Yu Guang Pu Fen Xi. 2010 Oct;30(10):2724-8. Guang Pu Xue Yu Guang Pu Fen Xi. 2010. PMID: 21137408 Chinese.
Cited by
-
Navigating the Functional Landscape of Transcription Factors via Non-Negative Tensor Factorization Analysis of MEDLINE Abstracts.Front Bioeng Biotechnol. 2017 Aug 28;5:48. doi: 10.3389/fbioe.2017.00048. eCollection 2017. Front Bioeng Biotechnol. 2017. PMID: 28894735 Free PMC article.
-
Orthogonal joint sparse NMF for microarray data analysis.J Math Biol. 2019 Jul;79(1):223-247. doi: 10.1007/s00285-019-01355-2. Epub 2019 Apr 19. J Math Biol. 2019. PMID: 31004215
References
-
- Shalon D, Smith SJ, Brown PO. A DNA microarray system for analyzing complex DNA samples using two-color fluorescent probe hybridization. Genome Research. 1996;6(7):639–645. - PubMed
-
- Granzow M, Berrar D, Dubitzky W, Schuste A, Azuaje F, Eils R. Tumor classification by gene expression profiling: comparison and validation of five clustering methods. ACM SIGBIO Newsletter. 2001;21:16–22.
-
- Wang L, Chu F, Xie W. Accurate cancer classification using expressions of very few genes. IEEE/ACM Transactions on Computational Biology and Bioinformatics. 2007;4(1):40–53. - PubMed
-
- Furey TS, Cristianini N, Duffy N, Bednarski DW, Schummer M, Haussler D. Support vector machine classification and validation of cancer tissue samples using microarray expression data. Bioinformatics. 2000;16(10):906–914. - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

