A comprehensive assessment of N-terminal signal peptides prediction methods
- PMID: 19958512
- PMCID: PMC2788353
- DOI: 10.1186/1471-2105-10-S15-S2
A comprehensive assessment of N-terminal signal peptides prediction methods
Abstract
Background: Amino-terminal signal peptides (SPs) are short regions that guide the targeting of secretory proteins to the correct subcellular compartments in the cell. They are cleaved off upon the passenger protein reaching its destination. The explosive growth in sequencing technologies has led to the deposition of vast numbers of protein sequences necessitating rapid functional annotation techniques, with subcellular localization being a key feature. Of the myriad software prediction tools developed to automate the task of assigning the SP cleavage site of these new sequences, we review here, the performance and reliability of commonly used SP prediction tools.
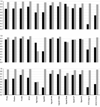
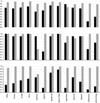
Results: The available signal peptide data has been manually curated and organized into three datasets representing eukaryotes, Gram-positive and Gram-negative bacteria. These datasets are used to evaluate thirteen prediction tools that are publicly available. SignalP (both the HMM and ANN versions) maintains consistency and achieves the best overall accuracy in all three benchmarking experiments, ranging from 0.872 to 0.914 although other prediction tools are narrowing the performance gap.
Conclusion: The majority of the tools evaluated in this study encounter no difficulty in discriminating between secretory and non-secretory proteins. The challenge clearly remains with pinpointing the correct SP cleavage site. The composite scoring schemes employed by SignalP may help to explain its accuracy. Prediction task is divided into a number of separate steps, thus allowing each score to tackle a particular aspect of the prediction.
Figures
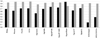
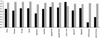
Similar articles
-
Flanking signal and mature peptide residues influence signal peptide cleavage.BMC Bioinformatics. 2008 Dec 12;9 Suppl 12(Suppl 12):S15. doi: 10.1186/1471-2105-9-S12-S15. BMC Bioinformatics. 2008. PMID: 19091014 Free PMC article.
-
Predicting Secretory Proteins with SignalP.Methods Mol Biol. 2017;1611:59-73. doi: 10.1007/978-1-4939-7015-5_6. Methods Mol Biol. 2017. PMID: 28451972
-
Signal peptide prediction based on analysis of experimentally verified cleavage sites.Protein Sci. 2004 Oct;13(10):2819-24. doi: 10.1110/ps.04682504. Epub 2004 Aug 31. Protein Sci. 2004. PMID: 15340161 Free PMC article.
-
Architecture, function and prediction of long signal peptides.Brief Bioinform. 2009 Sep;10(5):569-78. doi: 10.1093/bib/bbp030. Epub 2009 Jun 17. Brief Bioinform. 2009. PMID: 19535397 Review.
-
Machine learning approaches for the prediction of signal peptides and other protein sorting signals.Protein Eng. 1999 Jan;12(1):3-9. doi: 10.1093/protein/12.1.3. Protein Eng. 1999. PMID: 10065704 Review.
Cited by
-
Signatures of positive selection in Toll-like receptor (TLR) genes in mammals.BMC Evol Biol. 2011 Dec 20;11:368. doi: 10.1186/1471-2148-11-368. BMC Evol Biol. 2011. PMID: 22185391 Free PMC article.
-
Towards a career in bioinformatics.BMC Bioinformatics. 2009 Dec 3;10 Suppl 15(Suppl 15):S1. doi: 10.1186/1471-2105-10-S15-S1. BMC Bioinformatics. 2009. PMID: 19958508 Free PMC article.
-
Experimental annotation of post-translational features and translated coding regions in the pathogen Salmonella Typhimurium.BMC Genomics. 2011 Aug 25;12:433. doi: 10.1186/1471-2164-12-433. BMC Genomics. 2011. PMID: 21867535 Free PMC article.
-
Comprehensive Transcriptome Analysis Reveals Genome-Wide Changes Associated with Endoplasmic Reticulum (ER) Stress in Potato (Solanum tuberosum L.).Int J Mol Sci. 2022 Nov 9;23(22):13795. doi: 10.3390/ijms232213795. Int J Mol Sci. 2022. PMID: 36430273 Free PMC article.
-
The venom-gland transcriptome of the eastern coral snake (Micrurus fulvius) reveals high venom complexity in the intragenomic evolution of venoms.BMC Genomics. 2013 Aug 2;14:531. doi: 10.1186/1471-2164-14-531. BMC Genomics. 2013. PMID: 23915248 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

