Differential contribution of chemotaxis and substrate restriction to segregation of immature and mature thymocytes
- PMID: 19962328
- PMCID: PMC4106268
- DOI: 10.1016/j.immuni.2009.09.020
Differential contribution of chemotaxis and substrate restriction to segregation of immature and mature thymocytes
Abstract
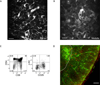
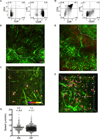
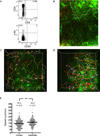
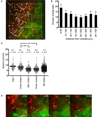
T cell development requires sequential localization of thymocyte subsets to distinct thymic microenvironments. To address mechanisms governing this segregation, we used two-photon microscopy to visualize migration of purified thymocyte subsets in defined microenvironments within thymic slices. Double-negative (CD4(-)8(-)) and double-positive (CD4(+)8(+)) thymocytes were confined to cortex where they moved slowly without directional bias. DP cells accumulated and migrated more rapidly in a specialized inner-cortical microenvironment, but were unable to migrate on medullary substrates. In contrast, CD4 single positive (SP) thymocytes migrated directionally toward the medulla, where they accumulated and moved very rapidly. Our results revealed a requisite two-step process governing CD4 SP cell medullary localization: the chemokine receptor CCR7 mediated chemotaxis of CD4 SP cells towards medulla, whereas a distinct pertussis-toxin sensitive pathway was required for medullary entry. These findings suggest that developmentally regulated responses to both chemotactic signals and specific migratory substrates guide thymocytes to specific locations in the thymus.
Copyright 2009 Elsevier Inc. All rights reserved.
Figures

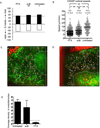
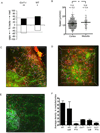
Comment in
-
Everything in its place.Immunity. 2009 Dec 18;31(6):856-8. doi: 10.1016/j.immuni.2009.11.005. Immunity. 2009. PMID: 20064445
References
-
- Anderson MS, Venanzi ES, Klein L, Chen Z, Berzins SP, Turley SJ, von Boehmer H, Bronson R, Dierich A, Benoist C, Mathis D. Projection of an immunological self shadow within the thymus by the aire protein. Science. 2002;298:1395–1401. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

