The appearance of pyrrolysine in tRNAHis guanylyltransferase by neutral evolution
- PMID: 19965368
- PMCID: PMC2795538
- DOI: 10.1073/pnas.0912072106
The appearance of pyrrolysine in tRNAHis guanylyltransferase by neutral evolution
Abstract
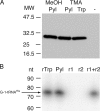
tRNA(His) guanylyltransferase (Thg1) post-transcriptionally adds a G (position -1) to the 5'-terminus of tRNA(His). The Methanosarcina acetivorans Thg1 (MaThg1) gene contains an in-frame TAG (amber) codon. Although a UAG codon typically directs translation termination, its presence in Methanosarcina mRNA may lead to pyrrolysine (Pyl) incorporation achieved by Pyl-tRNA(Pyl), the product of pyrrolysyl-tRNA synthetase. Sequencing of the MaThg1 gene and transcript confirmed the amber codon. Translation of MaThg1 mRNA led to a full-length, Pyl-containing, active enzyme as determined by immunoblotting, mass spectrometry, and biochemical analysis. The nature of the inserted amino acid at the position specified by UAG is not critical, as Pyl or Trp insertion yields active MaThg1 variants in M. acetivorans and equal amounts of full-length protein. These data suggest that Pyl insertion is akin to natural suppression and unlike the active stop codon reassignment that is required for selenocysteine insertion. Only three Pyl-containing proteins have been characterized previously, a set of methylamine methyltransferases in which Pyl is assumed to have specifically evolved to be a key active-site constituent. In contrast, Pyl in MaThg1 is a dispensable residue that appears to confer no selective advantage. Phylogenetic analysis suggests that Thg1 is becoming dispensable in the archaea, and furthermore supports the hypothesis that Pyl appeared in MaThg1 as the result of neutral evolution. This indicates that even the most unusual amino acid can play an ordinary role in proteins.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

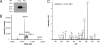

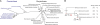
Similar articles
-
An aminoacyl-tRNA synthetase that specifically activates pyrrolysine.Proc Natl Acad Sci U S A. 2004 Aug 24;101(34):12450-4. doi: 10.1073/pnas.0405362101. Epub 2004 Aug 16. Proc Natl Acad Sci U S A. 2004. PMID: 15314242 Free PMC article.
-
Pyrrolysine and selenocysteine use dissimilar decoding strategies.J Biol Chem. 2005 May 27;280(21):20740-51. doi: 10.1074/jbc.M501458200. Epub 2005 Mar 22. J Biol Chem. 2005. PMID: 15788401
-
Characterization of a Methanosarcina acetivorans mutant unable to translate UAG as pyrrolysine.Mol Microbiol. 2006 Jan;59(1):56-66. doi: 10.1111/j.1365-2958.2005.04927.x. Mol Microbiol. 2006. PMID: 16359318
-
tRNAPyl: Structure, function, and applications.RNA Biol. 2018;15(4-5):441-452. doi: 10.1080/15476286.2017.1356561. Epub 2017 Sep 13. RNA Biol. 2018. PMID: 28837402 Free PMC article. Review.
-
Pyrrolysyl-tRNA synthetase: an ordinary enzyme but an outstanding genetic code expansion tool.Biochim Biophys Acta. 2014 Jun;1844(6):1059-70. doi: 10.1016/j.bbapap.2014.03.002. Epub 2014 Mar 12. Biochim Biophys Acta. 2014. PMID: 24631543 Free PMC article. Review.
Cited by
-
The requirement for the highly conserved G-1 residue of Saccharomyces cerevisiae tRNAHis can be circumvented by overexpression of tRNAHis and its synthetase.RNA. 2010 May;16(5):1068-77. doi: 10.1261/rna.2087510. Epub 2010 Apr 1. RNA. 2010. PMID: 20360392 Free PMC article.
-
Identification of distinct biological functions for four 3'-5' RNA polymerases.Nucleic Acids Res. 2016 Sep 30;44(17):8395-406. doi: 10.1093/nar/gkw681. Epub 2016 Aug 2. Nucleic Acids Res. 2016. PMID: 27484477 Free PMC article.
-
Change of tRNA identity leads to a divergent orthogonal histidyl-tRNA synthetase/tRNAHis pair.Nucleic Acids Res. 2011 Mar;39(6):2286-93. doi: 10.1093/nar/gkq1176. Epub 2010 Nov 17. Nucleic Acids Res. 2011. PMID: 21087993 Free PMC article.
-
Minimal requirements for reverse polymerization and tRNA repair by tRNAHis guanylyltransferase.RNA Biol. 2018;15(4-5):614-622. doi: 10.1080/15476286.2017.1372076. Epub 2017 Sep 29. RNA Biol. 2018. PMID: 28901837 Free PMC article.
-
In vitro substrate specificities of 3'-5' polymerases correlate with biological outcomes of tRNA 5'-editing reactions.FEBS Lett. 2015 Jul 22;589(16):2124-30. doi: 10.1016/j.febslet.2015.06.028. Epub 2015 Jul 2. FEBS Lett. 2015. PMID: 26143376 Free PMC article.
References
-
- Hao B, et al. A new UAG-encoded residue in the structure of a methanogen methyltransferase. Science. 2002;296:1462–1466. - PubMed
-
- Blight SK, et al. Direct charging of tRNACUA with pyrrolysine in vitro and in vivo. Nature. 2004;431:333–335. - PubMed
-
- Mahapatra A, et al. Characterization of a Methanosarcina acetivorans mutant unable to translate UAG as pyrrolysine. Mol Microbiol. 2006;59:56–66. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

