Crystal structure of the catalytic core of an RNA-polymerase ribozyme
- PMID: 19965478
- PMCID: PMC3978776
- DOI: 10.1126/science.1174676
Crystal structure of the catalytic core of an RNA-polymerase ribozyme
Abstract
Primordial organisms of the putative RNA world would have required polymerase ribozymes able to replicate RNA. Known ribozymes with polymerase activity best approximating that needed for RNA replication contain at their catalytic core the class I RNA ligase, an artificial ribozyme with a catalytic rate among the fastest of known ribozymes. Here we present the 3.0 angstrom crystal structure of this ligase. The architecture resembles a tripod, its three legs converging near the ligation junction. Interacting with this tripod scaffold through a series of 10 minor-groove interactions (including two A-minor triads) is the unpaired segment that contributes to and organizes the active site. A cytosine nucleobase and two backbone phosphates abut the ligation junction; their location suggests a model for catalysis resembling that of proteinaceous polymerases.
Figures


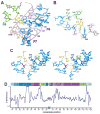

Similar articles
-
The structural basis of ribozyme-catalyzed RNA assembly.Science. 2007 Mar 16;315(5818):1549-53. doi: 10.1126/science.1136231. Science. 2007. PMID: 17363667
-
Crystal structure of an RNA polymerase ribozyme in complex with an antibody fragment.Philos Trans R Soc Lond B Biol Sci. 2011 Oct 27;366(1580):2918-28. doi: 10.1098/rstb.2011.0144. Philos Trans R Soc Lond B Biol Sci. 2011. PMID: 21930583 Free PMC article.
-
Characterization of the B6.61 polymerase ribozyme accessory domain.RNA. 2011 Mar;17(3):469-77. doi: 10.1261/rna.2495011. Epub 2011 Jan 11. RNA. 2011. PMID: 21224380 Free PMC article.
-
Ribozymes: structure and mechanism in RNA catalysis.Trends Biochem Sci. 1996 Jun;21(6):220-4. Trends Biochem Sci. 1996. PMID: 8744356 Review.
-
Structural Simplicity and Mechanistic Complexity in the Hammerhead Ribozyme.Prog Mol Biol Transl Sci. 2018;159:177-202. doi: 10.1016/bs.pmbts.2018.07.006. Epub 2018 Sep 17. Prog Mol Biol Transl Sci. 2018. PMID: 30340787 Review.
Cited by
-
Witnessing the structural evolution of an RNA enzyme.Elife. 2021 Sep 9;10:e71557. doi: 10.7554/eLife.71557. Elife. 2021. PMID: 34498588 Free PMC article.
-
Structure and catalytic activity of the SAM-utilizing ribozyme SAMURI.Nat Chem Biol. 2025 Jan 8. doi: 10.1038/s41589-024-01808-w. Online ahead of print. Nat Chem Biol. 2025. PMID: 39779902
-
Microfluidic compartmentalized directed evolution.Chem Biol. 2010 Jul 30;17(7):717-24. doi: 10.1016/j.chembiol.2010.05.021. Chem Biol. 2010. PMID: 20659684 Free PMC article.
-
In-ice evolution of RNA polymerase ribozyme activity.Nat Chem. 2013 Dec;5(12):1011-8. doi: 10.1038/nchem.1781. Epub 2013 Oct 20. Nat Chem. 2013. PMID: 24256864 Free PMC article.
-
Mapping L1 ligase ribozyme conformational switch.J Mol Biol. 2012 Oct 12;423(1):106-22. doi: 10.1016/j.jmb.2012.06.035. Epub 2012 Jul 3. J Mol Biol. 2012. PMID: 22771572 Free PMC article.
References
-
- Joyce GF, Orgel LE. In: The RNA World. Gesteland RF, Cech TR, Atkins JF, editors. Cold Spring Harbor Laboratory Press; Cold Spring Harbor, NY: 1999. pp. 49–77.
-
- Johnston WK, Unrau PJ, Lawrence MS, Glasner ME, Bartel DP. Science. 2001;292:1319. - PubMed
-
- Bartel DP, Szostak JW. Science. 1993;261:1411. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

