Prenatal exposure to environmental tobacco smoke alters gene expression in the developing murine hippocampus
- PMID: 19969065
- PMCID: PMC2844479
- DOI: 10.1016/j.reprotox.2009.12.001
Prenatal exposure to environmental tobacco smoke alters gene expression in the developing murine hippocampus
Abstract
Background: Little is known about the effects of passive smoke exposures on the developing brain.
Objective: The purpose of the current study was to identify changes in gene expression in the murine hippocampus as a consequence of in utero exposure to sidestream cigarette smoke (an experimental equivalent of environmental tobacco smoke (ETS)) at exposure levels that do not result in fetal growth inhibition.
Methods: A whole body smoke inhalation exposure system was utilized to deliver ETS to pregnant C57BL/6J mice for 6 h/day from gestational days 6-17 (gd 6-17) [for microarray] or gd 6-18.5 [for fetal phenotyping].
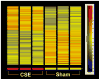
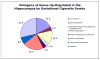
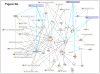
Results: There were no significant effects of ETS exposure on fetal phenotype. However, 61 "expressed" genes in the gd 18.5 fetal hippocampus were differentially regulated (up- or down-regulated by 1.5-fold or greater) by maternal exposure to ETS. Of these 61 genes, 25 genes were upregulated while 36 genes were down-regulated. A systems biology approach, including computational methodologies, identified cellular response pathways, and biological themes, underlying altered fetal programming of the embryonic hippocampus by in utero cigarette smoke exposure.
Conclusions: Results from the present study suggest that even in the absence of effects on fetal growth, prenatal smoke exposure can alter gene expression during the "early" period of hippocampal growth and may result in abnormal hippocampal morphology, connectivity, and function.
Copyright 2009 Elsevier Inc. All rights reserved.
Conflict of interest statement
The authors declare that there are no conflicts of interest.
Figures



References
-
- Centers for Disease Control and Prevention. Cigarette smoking among adults--United States, 2004. MMWR Morb Mortal Wkly Rep. 2005 Nov 11;54(44):1121–24. - PubMed
-
- Chen LH, Petitti DB. Case-control study of passive smoking and the risk of small-for-gestational-age at term. Am J Epidemiol. 1995 Jul 15;142(2):158–65. - PubMed
-
- Jordan TR, Price JH, Dake JA, Shah S. Adolescent exposure to and perceptions of environmental tobacco smoke. J Sch Health. 2005 May;75(5):178–186. - PubMed
-
- Rudatsikira EM, Knutsen SF, Job JS, et al. Exposure to environmental tobacco smoke in the nonsmoking population of Cambodia. Am J Prev Med. 2008 Jan;34(1):69–73. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

