Global MYCN transcription factor binding analysis in neuroblastoma reveals association with distinct E-box motifs and regions of DNA hypermethylation
- PMID: 19997598
- PMCID: PMC2781550
- DOI: 10.1371/journal.pone.0008154
Global MYCN transcription factor binding analysis in neuroblastoma reveals association with distinct E-box motifs and regions of DNA hypermethylation
Abstract
Background: Neuroblastoma, a cancer derived from precursor cells of the sympathetic nervous system, is a major cause of childhood cancer related deaths. The single most important prognostic indicator of poor clinical outcome in this disease is genomic amplification of MYCN, a member of a family of oncogenic transcription factors.
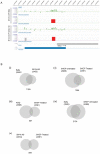
Methodology: We applied MYCN chromatin immunoprecipitation to microarrays (ChIP-chip) using MYCN amplified/non-amplified cell lines as well as a conditional knockdown cell line to determine the distribution of MYCN binding sites within all annotated promoter regions.
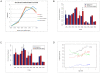
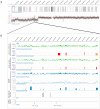
Conclusion: Assessment of E-box usage within consistently positive MYCN binding sites revealed a predominance for the CATGTG motif (p<0.0016), with significant enrichment of additional motifs CATTTG, CATCTG, CAACTG in the MYCN amplified state. For cell lines over-expressing MYCN, gene ontology analysis revealed enrichment for the binding of MYCN at promoter regions of numerous molecular functional groups including DNA helicases and mRNA transcriptional regulation. In order to evaluate MYCN binding with respect to other genomic features, we determined the methylation status of all annotated CpG islands and promoter sequences using methylated DNA immunoprecipitation (MeDIP). The integration of MYCN ChIP-chip and MeDIP data revealed a highly significant positive correlation between MYCN binding and DNA hypermethylation. This association was also detected in regions of hemizygous loss, indicating that the observed association occurs on the same homologue. In summary, these findings suggest that MYCN binding occurs more commonly at CATGTG as opposed to the classic CACGTG E-box motif, and that disease associated over expression of MYCN leads to aberrant binding to additional weaker affinity E-box motifs in neuroblastoma. The co-localization of MYCN binding and DNA hypermethylation further supports the dual role of MYCN, namely that of a classical transcription factor affecting the activity of individual genes, and that of a mediator of global chromatin structure.
Conflict of interest statement
Figures
References
-
- Mukherjee B, Morgenbesser SD, DePinho RA. Myc family oncoproteins function through a common pathway to transform normal cells in culture: cross-interference by Max and trans-acting dominant mutants. Genes Dev. 1992;6:1480–1492. - PubMed
-
- Brodeur GM, Seeger RC, Schwab M, Varmus HE, Bishop JM. Amplification of N-myc in untreated human neuroblastomas correlates with advanced disease stage. Science. 1984;224:1121–1124. - PubMed
-
- Alaminos M, Mora J, Cheung NK, Smith A, Qin J, et al. Genome-wide analysis of gene expression associated with MYCN in human neuroblastoma. Cancer Res. 2003;63:4538–4546. - PubMed
-
- Valentijn LJ, Koppen A, van Asperen R, Root HA, Haneveld F, et al. Inhibition of a new differentiation pathway in neuroblastoma by copy number defects of N-myc, Cdc42, and nm23 genes. Cancer Res. 2005;65:3136–3145. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

