Identification of an emerin-beta-catenin complex in the heart important for intercalated disc architecture and beta-catenin localisation
- PMID: 19997769
- PMCID: PMC11115513
- DOI: 10.1007/s00018-009-0219-8
Identification of an emerin-beta-catenin complex in the heart important for intercalated disc architecture and beta-catenin localisation
Abstract
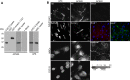
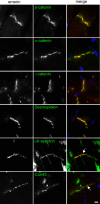
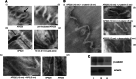
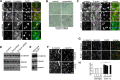
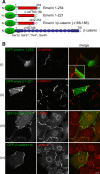
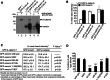
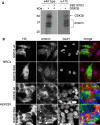
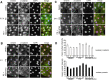
How mutations in the protein emerin lead to the cardiomyopathy associated with X-linked Emery-Dreifuss muscular dystrophy (X-EDMD) is unclear. We identified emerin at the adherens junction of the intercalated disc, where it co-localised with the catenin family of proteins. Emerin bound to wild type beta-catenin both in vivo and in vitro. Mutating the GSK3beta phosphorylation sites on beta-catenin abolished this binding. Wild type but not mutant forms of emerin associated with X-EDMD were able to reduce beta-catenin protein levels. Cardiomyocytes from emerin-null mice hearts exhibited erroneous beta-catenin distribution and intercalated disc architecture. Treatment of wild type cardiomyocytes with phenylephrine, which inactivates GSK3beta, redistributed emerin and beta-catenin. Emerin was identified as a direct target of GSK3beta activity since exogenous expression of GSK3beta reduced emerin levels at the nuclear envelope. We propose that perturbation to or total loss of the emerin-beta-catenin complex compromises both intercalated disc function and beta-catenin signalling in cardiomyocytes.
Figures
Similar articles
-
Emerin in health and disease.Semin Cell Dev Biol. 2014 May;29:95-106. doi: 10.1016/j.semcdb.2013.12.008. Epub 2013 Dec 21. Semin Cell Dev Biol. 2014. PMID: 24365856 Free PMC article. Review.
-
Attenuation of Wnt/β-catenin activity reverses enhanced generation of cardiomyocytes and cardiac defects caused by the loss of emerin.Hum Mol Genet. 2015 Feb 1;24(3):802-13. doi: 10.1093/hmg/ddu498. Epub 2014 Sep 30. Hum Mol Genet. 2015. PMID: 25274778
-
Emerin plays a crucial role in nuclear invagination and in the nuclear calcium transient.Sci Rep. 2017 Mar 14;7:44312. doi: 10.1038/srep44312. Sci Rep. 2017. PMID: 28290476 Free PMC article.
-
The cell cycle dependent mislocalisation of emerin may contribute to the Emery-Dreifuss muscular dystrophy phenotype.J Cell Sci. 2002 Jan 15;115(Pt 2):341-54. doi: 10.1242/jcs.115.2.341. J Cell Sci. 2002. PMID: 11839786
-
Molecular signatures of Emery-Dreifuss muscular dystrophy.Biochem Soc Trans. 2008 Dec;36(Pt 6):1354-8. doi: 10.1042/BST0361354. Biochem Soc Trans. 2008. PMID: 19021555 Review.
Cited by
-
Emerin in health and disease.Semin Cell Dev Biol. 2014 May;29:95-106. doi: 10.1016/j.semcdb.2013.12.008. Epub 2013 Dec 21. Semin Cell Dev Biol. 2014. PMID: 24365856 Free PMC article. Review.
-
Messages from the voices within: regulation of signaling by proteins of the nuclear lamina.Curr Opin Cell Biol. 2018 Jun;52:14-21. doi: 10.1016/j.ceb.2017.12.009. Epub 2018 Jan 4. Curr Opin Cell Biol. 2018. PMID: 29306725 Free PMC article. Review.
-
Novel nesprin-1 mutations associated with dilated cardiomyopathy cause nuclear envelope disruption and defects in myogenesis.Hum Mol Genet. 2017 Jun 15;26(12):2258-2276. doi: 10.1093/hmg/ddx116. Hum Mol Genet. 2017. PMID: 28398466 Free PMC article.
-
Activity-dependent synthesis of Emerin gates neuronal plasticity by regulating proteostasis.Cell Rep. 2025 Apr 22;44(4):115439. doi: 10.1016/j.celrep.2025.115439. Epub 2025 Apr 9. Cell Rep. 2025. PMID: 40208794 Free PMC article.
-
Cardiac manifestations and clinical management of X-linked Emery-Dreifuss muscular dystrophy: a case series.Eur Heart J Case Rep. 2023 Jan 11;7(1):ytad013. doi: 10.1093/ehjcr/ytad013. eCollection 2023 Jan. Eur Heart J Case Rep. 2023. PMID: 36727127 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

