Pathogenesis of ovarian cancer: clues from selected overexpressed genes
- PMID: 20001801
- PMCID: PMC2821471
- DOI: 10.2217/fon.09.126
Pathogenesis of ovarian cancer: clues from selected overexpressed genes
Abstract
Ovarian cancer is the most malignant gynecologic neoplasm. Although new chemotherapeutic agents have improved patients' 5-year survival rate, the overall mortality of ovarian cancer has remained largely unchanged in the past several decades. The main reason for the lack of success in effectively treating ovarian cancer is our limited understanding of its etiology and the very few molecular diagnostic markers and therapeutic targets known so far. Identification and characterization of ovarian cancer-associated genes are fundamental for unveiling the pathogenesis of its initiation and progression, especially the development of recurrent diseases. As there are a vast number of genes for which molecular genetic changes and aberrant gene expression have been reported in ovarian cancer, this review will only focus on summarizing those exemplified genes that have been demonstrated to have biological functions in promoting ovarian cancer development and potential clinical significance. The genes to be discussed include nuclear proteins (Notch3, HBXAP [Rsf-1], NAC1 and NFkappaB), cytoplasmic proteins (fatty acid synthase and apolipoprotein E) and cell surface/secretory proteins (mucin-4, mesothelin, claudin, HLA-G, kallikrein and folate receptor and osteopontin). Since the study of ovarian cancer-associated genes is complicated by several factors unique to ovarian cancer, we will also present our views on the limitations and challenges of current ovarian cancer research.
Figures


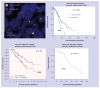
References
-
- Nakayama K, Nakayama N, Jinawath N, et al. Amplicon profiles in ovarian serous carcinomas. Int J Cancer. 2007;120:2613–2617. - PubMed
-
▪ Describes the genome-wide DNA copy number changes in serous ovarian cancer using purified tumor samples.
-
- Sturgeon CM, Duffy MJ, Stenman UH, et al. National Academy of Clinical Biochemistry laboratory medicine practice guidelines for use of tumor markers in testicular, prostate, colorectal, breast, and ovarian cancers. Clin Chem. 2008;54:E11–79. - PubMed
-
▪▪ A comprehensive overview of protein biomarkers in ovarian cancer.
-
- Shih IM, Sokoll LJ, Chan DW. Tumor markers in ovarian cancer. In: Diamandis EP, Fritsche HA, Lilja H, Chan DW, Schwartz MK, editors. Tumor markers physiology, pathobiology, technology and clinical applications. AACC Press; PA, USA: 2002. pp. 239–252.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous
