DescFold: a web server for protein fold recognition
- PMID: 20003426
- PMCID: PMC2803855
- DOI: 10.1186/1471-2105-10-416
DescFold: a web server for protein fold recognition
Abstract
Background: Machine learning-based methods have been proven to be powerful in developing new fold recognition tools. In our previous work [Zhang, Kochhar and Grigorov (2005) Protein Science, 14: 431-444], a machine learning-based method called DescFold was established by using Support Vector Machines (SVMs) to combine the following four descriptors: a profile-sequence-alignment-based descriptor using Psi-blast e-values and bit scores, a sequence-profile-alignment-based descriptor using Rps-blast e-values and bit scores, a descriptor based on secondary structure element alignment (SSEA), and a descriptor based on the occurrence of PROSITE functional motifs. In this work, we focus on the improvement of DescFold by incorporating more powerful descriptors and setting up a user-friendly web server.
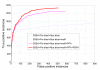
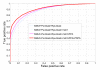
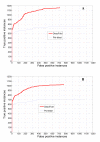
Results: In seeking more powerful descriptors, the profile-profile alignment score generated from the COMPASS algorithm was first considered as a new descriptor (i.e., PPA). When considering a profile-profile alignment between two proteins in the context of fold recognition, one protein is regarded as a template (i.e., its 3D structure is known). Instead of a sequence profile derived from a Psi-blast search, a structure-seeded profile for the template protein was generated by searching its structural neighbors with the assistance of the TM-align structural alignment algorithm. Moreover, the COMPASS algorithm was used again to derive a profile-structural-profile-alignment-based descriptor (i.e., PSPA). We trained and tested the new DescFold in a total of 1,835 highly diverse proteins extracted from the SCOP 1.73 version. When the PPA and PSPA descriptors were introduced, the new DescFold boosts the performance of fold recognition substantially. Using the SCOP_1.73_40% dataset as the fold library, the DescFold web server based on the trained SVM models was further constructed. To provide a large-scale test for the new DescFold, a stringent test set of 1,866 proteins were selected from the SCOP 1.75 version. At a less than 5% false positive rate control, the new DescFold is able to correctly recognize structural homologs at the fold level for nearly 46% test proteins. Additionally, we also benchmarked the DescFold method against several well-established fold recognition algorithms through the LiveBench targets and Lindahl dataset.
Conclusions: The new DescFold method was intensively benchmarked to have very competitive performance compared with some well-established fold recognition methods, suggesting that it can serve as a useful tool to assist in template-based protein structure prediction. The DescFold server is freely accessible at http://202.112.170.199/DescFold/index.html.
Figures

Similar articles
-
Descriptor-based protein remote homology identification.Protein Sci. 2005 Feb;14(2):431-44. doi: 10.1110/ps.041035505. Epub 2005 Jan 4. Protein Sci. 2005. PMID: 15632283 Free PMC article.
-
SVM-Fold: a tool for discriminative multi-class protein fold and superfamily recognition.BMC Bioinformatics. 2007 May 22;8 Suppl 4(Suppl 4):S2. doi: 10.1186/1471-2105-8-S4-S2. BMC Bioinformatics. 2007. PMID: 17570145 Free PMC article.
-
TIM-Finder: a new method for identifying TIM-barrel proteins.BMC Struct Biol. 2009 Dec 14;9:73. doi: 10.1186/1472-6807-9-73. BMC Struct Biol. 2009. PMID: 20003393 Free PMC article.
-
Comparison of proteins based on segments structural similarity.Acta Biochim Pol. 2004;51(1):161-72. Acta Biochim Pol. 2004. PMID: 15094837 Review.
-
A simple recipe for the non-expert bioinformaticist for building experimentally-testable hypotheses for proteins with no known homologs.J Struct Funct Genomics. 2012 Dec;13(4):185-200. doi: 10.1007/s10969-012-9141-7. Epub 2012 Sep 7. J Struct Funct Genomics. 2012. PMID: 22956349 Review.
Cited by
-
Outer membrane proteins can be simply identified using secondary structure element alignment.BMC Bioinformatics. 2011 Mar 17;12:76. doi: 10.1186/1471-2105-12-76. BMC Bioinformatics. 2011. PMID: 21414186 Free PMC article.
-
ProFold: Protein Fold Classification with Additional Structural Features and a Novel Ensemble Classifier.Biomed Res Int. 2016;2016:6802832. doi: 10.1155/2016/6802832. Epub 2016 Aug 28. Biomed Res Int. 2016. PMID: 27660761 Free PMC article.
-
SVM-SulfoSite: A support vector machine based predictor for sulfenylation sites.Sci Rep. 2018 Jul 26;8(1):11288. doi: 10.1038/s41598-018-29126-x. Sci Rep. 2018. PMID: 30050050 Free PMC article.
-
Incorporation of local structural preference potential improves fold recognition.PLoS One. 2011 Feb 18;6(2):e17215. doi: 10.1371/journal.pone.0017215. PLoS One. 2011. PMID: 21365008 Free PMC article.
-
EPuL: An Enhanced Positive-Unlabeled Learning Algorithm for the Prediction of Pupylation Sites.Molecules. 2017 Sep 5;22(9):1463. doi: 10.3390/molecules22091463. Molecules. 2017. PMID: 28872627 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

