Estimates of linkage disequilibrium and effective population size in rainbow trout
- PMID: 20003428
- PMCID: PMC2800115
- DOI: 10.1186/1471-2156-10-83
Estimates of linkage disequilibrium and effective population size in rainbow trout
Abstract
Background: The use of molecular genetic technologies for broodstock management and selective breeding of aquaculture species is becoming increasingly more common with the continued development of genome tools and reagents. Several laboratories have produced genetic maps for rainbow trout to aid in the identification of loci affecting phenotypes of interest. These maps have resulted in the identification of many quantitative/qualitative trait loci affecting phenotypic variation in traits associated with albinism, disease resistance, temperature tolerance, sex determination, embryonic development rate, spawning date, condition factor and growth. Unfortunately, the elucidation of the precise allelic variation and/or genes underlying phenotypic diversity has yet to be achieved in this species having low marker densities and lacking a whole genome reference sequence. Experimental designs which integrate segregation analyses with linkage disequilibrium (LD) approaches facilitate the discovery of genes affecting important traits. To date the extent of LD has been characterized for humans and several agriculturally important livestock species but not for rainbow trout.
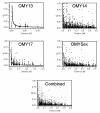
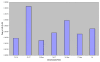
Results: We observed that the level of LD between syntenic loci decayed rapidly at distances greater than 2 cM which is similar to observations of LD in other agriculturally important species including cattle, sheep, pigs and chickens. However, in some cases significant LD was also observed up to 50 cM. Our estimate of effective population size based on genome wide estimates of LD for the NCCCWA broodstock population was 145, indicating that this population will respond well to high selection intensity. However, the range of effective population size based on individual chromosomes was 75.51 - 203.35, possibly indicating that suites of genes on each chromosome are disproportionately under selection pressures.
Conclusions: Our results indicate that large numbers of markers, more than are currently available for this species, will be required to enable the use of genome-wide integrated mapping approaches aimed at identifying genes of interest in rainbow trout.
Figures
Similar articles
-
A first generation integrated map of the rainbow trout genome.BMC Genomics. 2011 Apr 7;12:180. doi: 10.1186/1471-2164-12-180. BMC Genomics. 2011. PMID: 21473775 Free PMC article.
-
A second generation genetic map for rainbow trout (Oncorhynchus mykiss).BMC Genet. 2008 Nov 19;9:74. doi: 10.1186/1471-2156-9-74. BMC Genet. 2008. PMID: 19019240 Free PMC article.
-
QTL affecting stress response to crowding in a rainbow trout broodstock population.BMC Genet. 2012 Nov 7;13:97. doi: 10.1186/1471-2156-13-97. BMC Genet. 2012. PMID: 23134666 Free PMC article.
-
A first generation BAC-based physical map of the rainbow trout genome.BMC Genomics. 2009 Oct 8;10:462. doi: 10.1186/1471-2164-10-462. BMC Genomics. 2009. PMID: 19814815 Free PMC article.
-
A type I and type II microsatellite linkage map of rainbow trout (Oncorhynchus mykiss) with presumptive coverage of all chromosome arms.BMC Genomics. 2006 Nov 30;7:302. doi: 10.1186/1471-2164-7-302. BMC Genomics. 2006. PMID: 17137492 Free PMC article.
Cited by
-
Association mapping of disease resistance traits in rainbow trout using restriction site associated DNA sequencing.G3 (Bethesda). 2014 Oct 28;4(12):2473-81. doi: 10.1534/g3.114.014621. G3 (Bethesda). 2014. PMID: 25354781 Free PMC article.
-
Linkage disequilibrium, persistence of phase and effective population size estimates in Hereford and Braford cattle.BMC Genet. 2016 Feb 1;17:32. doi: 10.1186/s12863-016-0339-8. BMC Genet. 2016. PMID: 26832943 Free PMC article.
-
The Global Durum Wheat Panel (GDP): An International Platform to Identify and Exchange Beneficial Alleles.Front Plant Sci. 2020 Dec 21;11:569905. doi: 10.3389/fpls.2020.569905. eCollection 2020. Front Plant Sci. 2020. PMID: 33408724 Free PMC article.
-
Genetic Diversity within a Collection of Italian Maize Inbred Lines: A Resource for Maize Genomics and Breeding.Plants (Basel). 2024 Jan 23;13(3):336. doi: 10.3390/plants13030336. Plants (Basel). 2024. PMID: 38337869 Free PMC article.
-
A first generation integrated map of the rainbow trout genome.BMC Genomics. 2011 Apr 7;12:180. doi: 10.1186/1471-2164-12-180. BMC Genomics. 2011. PMID: 21473775 Free PMC article.
References
-
- Liu ZJ, Ed. Aquaculture Genome Technologies. Ames, Iowa: Blackwell Publishing; 2007.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

