Global analysis of short RNAs reveals widespread promoter-proximal stalling and arrest of Pol II in Drosophila
- PMID: 20007866
- PMCID: PMC3435875
- DOI: 10.1126/science.1181421
Global analysis of short RNAs reveals widespread promoter-proximal stalling and arrest of Pol II in Drosophila
Abstract
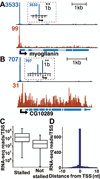
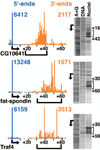
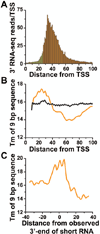
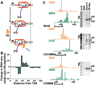
Emerging evidence indicates that gene expression in higher organisms is regulated by RNA polymerase II stalling during early transcription elongation. To probe the mechanisms responsible for this regulation, we developed methods to isolate and characterize short RNAs derived from stalled RNA polymerase II in Drosophila cells. Significant levels of these short RNAs were generated from more than one-third of all genes, indicating that promoter-proximal stalling is a general feature of early polymerase elongation. Nucleotide composition of the initially transcribed sequence played an important role in promoting transcriptional stalling by rendering polymerase elongation complexes highly susceptible to backtracking and arrest. These results indicate that the intrinsic efficiency of early elongation can greatly affect gene expression.
Figures
References
-
- Krumm A, Hickey LB, Groudine M. Genes Dev. 1995;9:559. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

