Correlation of Dehalococcoides 16S rRNA and chloroethene-reductive dehalogenase genes with geochemical conditions in chloroethene-contaminated groundwater
- PMID: 20008170
- PMCID: PMC2813033
- DOI: 10.1128/AEM.01482-09
Correlation of Dehalococcoides 16S rRNA and chloroethene-reductive dehalogenase genes with geochemical conditions in chloroethene-contaminated groundwater
Abstract
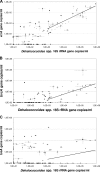
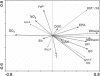
Quantitative analysis of genes that code for Dehalococcoides 16S rRNA and chloroethene-reductive dehalogenases TceA, VcrA, and BvcA was done on groundwater sampled from 150 monitoring wells spread over 11 chlorinated ethene polluted European locations. Redundancy analysis was used to relate molecular data to geochemical conditions. Dehalococcoides 16S rRNA- and vinyl chloride (VC)-reductase genes were present at all tested locations in concentrations up to 10(6) gene copies per ml of groundwater. However, differences between and also within locations were observed. Variation in Dehalococcoides 16S rRNA gene copy numbers were most strongly correlated to dissolved organic carbon concentration in groundwater and to conditions appropriate for biodegradation of chlorinated ethenes (U.S. Environmental Protection Agency score). In contrast, vcrA gene copy numbers correlated most significantly to VC and chlorinated ethene concentrations. Interestingly, bvcA and especially tceA were more correlated with oxidizing conditions. In groundwater microcosms, dechlorination of 1 mM VC was correlated to an increase of vcrA and/or bvcA gene copies by 2 to 4 orders of magnitude. Interestingly, in 34% of the monitoring wells and in 40% of the active microcosms, the amount of individual VC-reductase gene copies exceeded that of Dehalococcoides 16S rRNA gene copies. It is concluded that the geographical distribution of the genes was not homogeneous, depending on the geochemical conditions, whereby tceA and bvcA correlated to more oxidized conditions than Dehalococcoides 16S rRNA and vcrA. Because the variation in VC-reductase gene numbers was not directly correlated to variation in Dehalococcoides spp., VC-reductase genes are better monitoring parameters for VC dechlorination capacity than Dehalococcoides spp.
Figures
Similar articles
-
Quantifying genes and transcripts to assess the in situ physiology of "Dehalococcoides" spp. in a trichloroethene-contaminated groundwater site.Appl Environ Microbiol. 2008 May;74(9):2728-39. doi: 10.1128/AEM.02199-07. Epub 2008 Mar 7. Appl Environ Microbiol. 2008. PMID: 18326677 Free PMC article.
-
Monitoring abundance and expression of "Dehalococcoides" species chloroethene-reductive dehalogenases in a tetrachloroethene-dechlorinating flow column.Appl Environ Microbiol. 2008 Sep;74(18):5695-703. doi: 10.1128/AEM.00926-08. Epub 2008 Aug 1. Appl Environ Microbiol. 2008. PMID: 18676701 Free PMC article.
-
Normalized Quantitative PCR Measurements as Predictors for Ethene Formation at Sites Impacted with Chlorinated Ethenes.Environ Sci Technol. 2018 Nov 20;52(22):13410-13420. doi: 10.1021/acs.est.8b04373. Epub 2018 Nov 8. Environ Sci Technol. 2018. PMID: 30365883 Free PMC article.
-
The little bacteria that can - diversity, genomics and ecophysiology of 'Dehalococcoides' spp. in contaminated environments.Microb Biotechnol. 2010 Jul;3(4):389-402. doi: 10.1111/j.1751-7915.2009.00147.x. Epub 2009 Sep 4. Microb Biotechnol. 2010. PMID: 21255338 Free PMC article. Review.
-
Effect of temperature on microbial reductive dehalogenation of chlorinated ethenes: a review.FEMS Microbiol Ecol. 2022 Sep 1;98(9):fiac081. doi: 10.1093/femsec/fiac081. FEMS Microbiol Ecol. 2022. PMID: 35810002 Review.
Cited by
-
Comparative genomics of two newly isolated Dehalococcoides strains and an enrichment using a genus microarray.ISME J. 2011 Jun;5(6):1014-24. doi: 10.1038/ismej.2010.202. Epub 2011 Jan 13. ISME J. 2011. PMID: 21228894 Free PMC article.
-
Chlorinated Electron Acceptor Abundance Drives Selection of Dehalococcoides mccartyi (D. mccartyi) Strains in Dechlorinating Enrichment Cultures and Groundwater Environments.Front Microbiol. 2018 May 17;9:812. doi: 10.3389/fmicb.2018.00812. eCollection 2018. Front Microbiol. 2018. PMID: 29867784 Free PMC article.
-
Selective enrichment yields robust ethene-producing dechlorinating cultures from microcosms stalled at cis-dichloroethene.PLoS One. 2014 Jun 20;9(6):e100654. doi: 10.1371/journal.pone.0100654. eCollection 2014. PLoS One. 2014. PMID: 24950250 Free PMC article.
-
Quantitative PCR for tracking the megaplasmid-borne biodegradation potential of a model sphingomonad.Appl Environ Microbiol. 2012 Jun;78(12):4493-6. doi: 10.1128/AEM.00715-12. Epub 2012 Apr 6. Appl Environ Microbiol. 2012. PMID: 22492441 Free PMC article.
-
Degradation Characteristics and Remediation Ability of Contaminated Soils by Using β-HCH Degrading Bacteria.Int J Environ Res Public Health. 2023 Feb 4;20(4):2767. doi: 10.3390/ijerph20042767. Int J Environ Res Public Health. 2023. PMID: 36833464 Free PMC article.
References
-
- Amos, B. K., K. M. Ritalahti, C. Cruz-Garcia, E. Padilla-Crespo, and F. E. Löffler. 2008. Oxygen effect on Dehalococcoides viability and biomarker quantification. Environ. Sci. Technol. 42:5718-5726. - PubMed
-
- Behrens, S., M. F. Azizian, P. J. McMurdie, A. Sabalowsky, M. E. Dolan, L. Semprini, and A. M. Spormann. 2008. Monitoring abundance and expression of “Dehalococcoides” species chloroethene-reductive dehalogenases in a tetrachloroethene-dechlorinating flow column. Appl. Environ. Microbiol. 74:5695-5703. - PMC - PubMed
-
- Cupples, A. M. 2008. Real-time PCR quantification of Dehalococcoides populations: methods and applications. J. Microbiol. Methods 72:1-11. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

