Selections that optimize RNA display in the yeast three-hybrid system
- PMID: 20008486
- PMCID: PMC2811653
- DOI: 10.1261/rna.1880410
Selections that optimize RNA display in the yeast three-hybrid system
Abstract
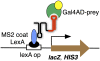
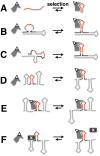
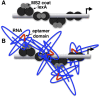
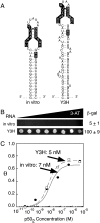
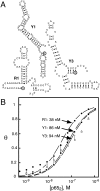
The yeast three-hybrid system (Y3H) is a powerful tool to select or confirm RNA-protein interactions. Target protein recognition of an RNA insert within a test transcript depends on at least three factors: intrinsic protein affinity for the properly folded insert, retention of RNA insert tertiary structure within a longer RNA transcript, and accessibility of the RNA insert to the target protein. Y3H reporter gene readout reflects the combination of these factors. Here, we discuss RNA insert tertiary structure and accessibility in the Y3H as "RNA display." We review evidence that RNA display can sometimes be optimized during Y3H selections that do not increase the intrinsic affinity of an RNA insert for a target protein. This situation is more likely when a library of RNA inserts and heterogeneous flanking sequences is subjected to selection, and is less likely when point mutations are targeted to the insert in a fixed context. An RNA display vector with enhanced modularity has been developed to minimize sequence context effects in the Y3H.
Figures
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
