Expression of BfrH, a putative siderophore receptor of Bordetella bronchiseptica, is regulated by iron, Fur1, and the extracellular function sigma factor EcfI
- PMID: 20008538
- PMCID: PMC2825947
- DOI: 10.1128/IAI.00961-09
Expression of BfrH, a putative siderophore receptor of Bordetella bronchiseptica, is regulated by iron, Fur1, and the extracellular function sigma factor EcfI
Abstract
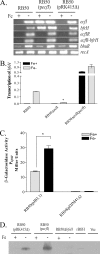
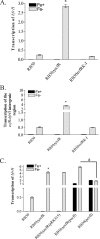
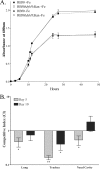
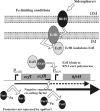
Iron (Fe) in soluble elemental form is found in the tissues and fluids of animals at concentrations insufficient for sustaining growth of bacteria. Consequently, to promote colonization and persistence, pathogenic bacteria evolved a myriad of scavenging mechanisms to acquire Fe from the host. Bordetella bronchiseptica, the etiologic agent of upper respiratory infections in a wide range of mammalian hosts, expresses a number of proteins for acquisition of Fe. Using proteomic and genomic approaches, three Fe-regulated genes were identified in the bordetellae: bfrH, a gene encoding a putative siderophore receptor; ecfI, a gene encoding a putative extracellular function (ECF) sigma factor; and ecfR, a gene encoding a putative EcfI modulator. All three genes are highly conserved in B. pertussis, B. parapertussis, and B. avium. Genetic analysis revealed that transcription of bfrH was coregulated by ecfI, ecfR, and fur1, one of two fur homologues carried by B. bronchiseptica. Overexpression of ecfI decoupled bfrH from Fe-dependent regulation. In contrast, expression of bfrH was significantly reduced in an ecfI deletion mutant. Deletion of ecfR, however, was correlated with a significant increase in expression of bfrH, due in part to a cis-acting nucleotide sequence within ecfR which likely reduces the frequency of readthrough transcription of bfrH from the Fe-dependent ecfIR promoter. Using a murine competition infection model, bfrH was shown to be required for optimal virulence of B. bronchiseptica. These experiments revealed ecfIR-bfrH as a locus encoding a new member of the growing family of Fe and ECF sigma factor-modulated regulons in the bordetellae.
Figures



Similar articles
-
Essential role of the iron-regulated outer membrane receptor FauA in alcaligin siderophore-mediated iron uptake in Bordetella species.J Bacteriol. 1999 Oct;181(19):5958-66. doi: 10.1128/JB.181.19.5958-5966.1999. J Bacteriol. 1999. PMID: 10498707 Free PMC article.
-
An iron-regulated outer-membrane protein specific to Bordetella bronchiseptica and homologous to ferric siderophore receptors.Microbiology (Reading). 1997 Jan;143 ( Pt 1):135-145. doi: 10.1099/00221287-143-1-135. Microbiology (Reading). 1997. PMID: 9025287
-
Interspecies variations in Bordetella catecholamine receptor gene regulation and function.Infect Immun. 2015 Dec;83(12):4639-52. doi: 10.1128/IAI.00787-15. Epub 2015 Sep 14. Infect Immun. 2015. PMID: 26371128 Free PMC article.
-
Environmental sensing mechanisms in Bordetella.Adv Microb Physiol. 2001;44:141-81. doi: 10.1016/s0065-2911(01)44013-6. Adv Microb Physiol. 2001. PMID: 11407112 Review.
-
Extracytoplasmic Function Sigma Factors Governing Production of the Primary Siderophores in Pathogenic Burkholderia Species.Front Microbiol. 2022 Feb 24;13:851011. doi: 10.3389/fmicb.2022.851011. eCollection 2022. Front Microbiol. 2022. PMID: 35283809 Free PMC article. Review.
Cited by
-
The Sigma Factor AsbI Is Required for the Expression of Acinetobactin Siderophore Transport Genes in Aeromonas salmonicida.Int J Mol Sci. 2023 Jun 2;24(11):9672. doi: 10.3390/ijms24119672. Int J Mol Sci. 2023. PMID: 37298622 Free PMC article.
-
Comparative Phosphoproteomics of Classical Bordetellae Elucidates the Potential Role of Serine, Threonine and Tyrosine Phosphorylation in Bordetella Biology and Virulence.Front Cell Infect Microbiol. 2021 Apr 13;11:660280. doi: 10.3389/fcimb.2021.660280. eCollection 2021. Front Cell Infect Microbiol. 2021. PMID: 33928046 Free PMC article.
-
Transcriptional profiling of the iron starvation response in Bordetella pertussis provides new insights into siderophore utilization and virulence gene expression.J Bacteriol. 2011 Sep;193(18):4798-812. doi: 10.1128/JB.05136-11. Epub 2011 Jul 8. J Bacteriol. 2011. PMID: 21742863 Free PMC article.
References
-
- Ahn, B. E., J. Cha, E. J. Lee, A. R. Han, C. J. Thompson, and J. H. Roe. 2006. Nur, a nickel-responsive regulator of the Fur family, regulates superoxide dismutases and nickel transport in Streptomyces coelicolor. Mol. Microbiol. 59:1848-1858. - PubMed
-
- Angerer, A., S. Enz, M. Ochs, and V. Braun. 1995. Transcriptional regulation of ferric citrate transport in Escherichia coli K-12. Fecl belongs to a new subfamily of sigma 70-type factors that respond to extracytoplasmic stimuli. Mol. Microbiol. 18:163-174. - PubMed
-
- Beall, B. 1998. Two iron-regulated putative ferric siderophore receptor genes in Bordetella bronchiseptica and Bordetella pertussis. Res. Microbiol. 149:189-201. - PubMed
-
- Beall, B., and T. Hoenes. 1997. An iron-regulated outer-membrane protein specific to Bordetella bronchiseptica and homologous to ferric siderophore receptors. Microbiology 143:135-145. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

